Analysis of 3k T cells from cancer#
import muon as mu
import numpy as np
import scanpy as sc
import scirpy as ir
from cycler import cycler
from matplotlib import cm as mpl_cm
from matplotlib import pyplot as plt
sc.set_figure_params(figsize=(4, 4))
sc.settings.verbosity = 2 # verbosity: errors (0), warnings (1), info (2), hints (3)
sc.logging.print_header()
scanpy==1.9.3 anndata==0.9.2 umap==0.5.3 numpy==1.24.4 scipy==1.11.1 pandas==2.0.3 scikit-learn==1.3.0 statsmodels==0.14.0 python-igraph==0.10.6 pynndescent==0.5.10
In this tutorial, we re-analyze single-cell TCR/RNA-seq data from Wu et al. [WMdA+20]. generated on the 10x Genomics platform. The original dataset consists of >140k T cells from 14 treatment-naive patients across four different types of cancer. For this tutorial, to speed up computations, we use a downsampled version of 3k cells.
Importing data#
Scirpy natively supports reading IR data from Cellranger (10x),
TraCeR (Smart-seq2)
or the AIRR rearrangement schema and provides helper functions to import other data types. We provide a dedicated tutorial on data loading with more details.
Warning
scirpy’s data structure has been updated in v0.13.0.
Previously, receptor data was expanded into columns of adata.obs, now they are stored as an awkward array in adata.obsm["airr"].
Moreover, we now use MuData to handle paired transcriptomics and AIRR data.
AnnData objects created with older versions of scirpy can be upgraded with scirpy.io.upgrade_schema() to be compatible with the latest version of scirpy.
Please check out
The release notes for details about the changes
The documentation about Scirpy’s data structure and
The example dataset for this tutorial ships with the scirpy package. We can conveniently load it from the datasets module:
mdata = ir.datasets.wu2020_3k()
mdata is a MuData object with two modalities: gex for gene expression and airr for immune-receptor data.
mdata
MuData object with n_obs × n_vars = 3000 × 30727
2 modalities
gex: 3000 x 30727
obs: 'cluster_orig', 'patient', 'sample', 'source'
uns: 'cluster_orig_colors'
obsm: 'X_umap_orig'
airr: 3000 x 0
obs: 'high_confidence', 'is_cell', 'clonotype_orig'
obsm: 'airr'Preprocessing Transcriptomics data#
Gene expression (GEX) data needs to be filtered and preprocessed as with any other single-cell dataset. For this tutorial, we perform minimal preprocessing for demonstration purposes. For a real dataset, we recommend following the scverse tutorials and the single-cell best practice book.
sc.pp.filter_genes(mdata["gex"], min_cells=10)
sc.pp.filter_cells(mdata["gex"], min_genes=100)
filtered out 18877 genes that are detected in less than 10 cells
sc.pp.normalize_per_cell(mdata["gex"])
sc.pp.log1p(mdata["gex"])
sc.pp.highly_variable_genes(mdata["gex"], flavor="cell_ranger", n_top_genes=5000)
sc.tl.pca(mdata["gex"])
sc.pp.neighbors(mdata["gex"])
normalizing by total count per cell
finished (0:00:00): normalized adata.X and added 'n_counts', counts per cell before normalization (adata.obs)
If you pass `n_top_genes`, all cutoffs are ignored.
extracting highly variable genes
finished (0:00:00)
computing PCA
on highly variable genes
with n_comps=50
finished (0:00:00)
computing neighbors
using 'X_pca' with n_pcs = 50
finished (0:00:01)
For the Wu2020 dataset, the authors already provide clusters and UMAP coordinates. Instead of performing clustering and cluster annotation ourselves, we will just use the provided data. The clustering and annotation methodology is described in their paper.
mdata["gex"].obsm["X_umap"] = mdata["gex"].obsm["X_umap_orig"]
mapping = {
"nan": "other",
"3.1-MT": "other",
"4.1-Trm": "CD4_Trm",
"4.2-RPL32": "CD4_RPL32",
"4.3-TCF7": "CD4_TCF7",
"4.4-FOS": "CD4_FOSS",
"4.5-IL6ST": "CD4_IL6ST",
"4.6a-Treg": "CD4_Treg",
"4.6b-Treg": "CD4_Treg",
"8.1-Teff": "CD8_Teff",
"8.2-Tem": "CD8_Tem",
"8.3a-Trm": "CD8_Trm",
"8.3b-Trm": "CD8_Trm",
"8.3c-Trm": "CD8_Trm",
"8.4-Chrom": "other",
"8.5-Mitosis": "other",
"8.6-KLRB1": "other",
}
mdata["gex"].obs["cluster"] = mdata["gex"].obs["cluster_orig"].map(mapping)
Now that we filtered obs and var of the GEX modality, we need to propagate those changes back to the MuData object.
mdata.update()
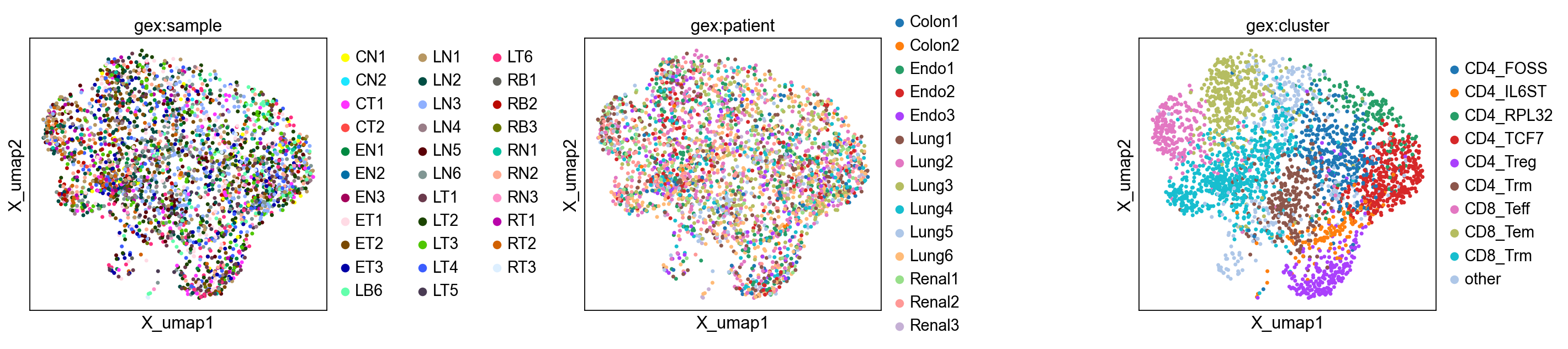
Let’s inspect the UMAP plots. The first three panels show the UMAP plot colored by sample, patient and cluster. We don’t observe any obvious clustering of samples or patients that could hint at batch effects.
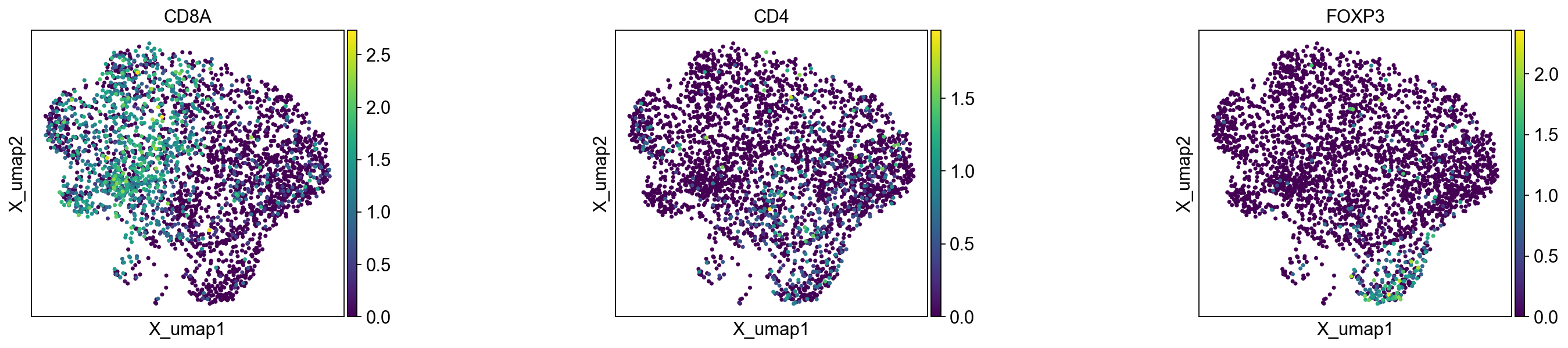
The last three panels show the UMAP colored by the T cell markers CD8, CD4, and FOXP3. We can confirm that the markers correspond to their respective cluster labels.
Creating chain indices#
As a first step, we need to create chain indices for the receptor data.
By importing IR data, essentially, for each cell a list of receptor chains is stored in adata.obsm["airr"].
The scirpy receptor model allows up to two pairs of chains per cell. This representation
requires separation of chains by locus into VJ and VDJ chains,
and (optionally) filtering non-productive chains.
The index_chains() function applies the scirpy receptor model by storing references to the
selected chains in adata.obsm["chain_indices"].
ir.pp.index_chains(mdata)
Filtering chains...
Indexing VJ chains...
Indexing VDJ chains...
build result array
TCR Quality Control#
While most of T cell receptors have exactly one pair of α and β chains, up to one third of T cells can have dual TCRs, i.e. two pairs of receptors originating from different alleles ([SB19]).
Using the scirpy.tl.chain_qc() function, we can add a summary
about the Immune cell-receptor compositions to adata.obs. We can visualize it using scirpy.pl.group_abundance().
Note
chain pairing
Orphan chain refers to cells that have either a single alpha or beta receptor chain.
Extra chain refers to cells that have a full alpha/beta receptor pair, and an additional chain.
Multichain refers to cells with more than two receptor pairs detected. These cells are likely doublets.
receptor type and receptor subtype
receptor_typerefers to a coarse classification intoBCRandTCR. Cells with bothBCRandTCRchains are labelled asambiguous.receptor_subtyperefers to a more specific classification into α/β, ɣ/δ, IG-λ, and IG-κ chain configurations.
For more details, see scirpy.tl.chain_qc().
ir.tl.chain_qc(mdata)
Stored result in `mdata.obs["airr:receptor_type"]`.
Stored result in `mdata.obs["airr:receptor_subtype"]`.
Stored result in `mdata.obs["airr:chain_pairing"]`.
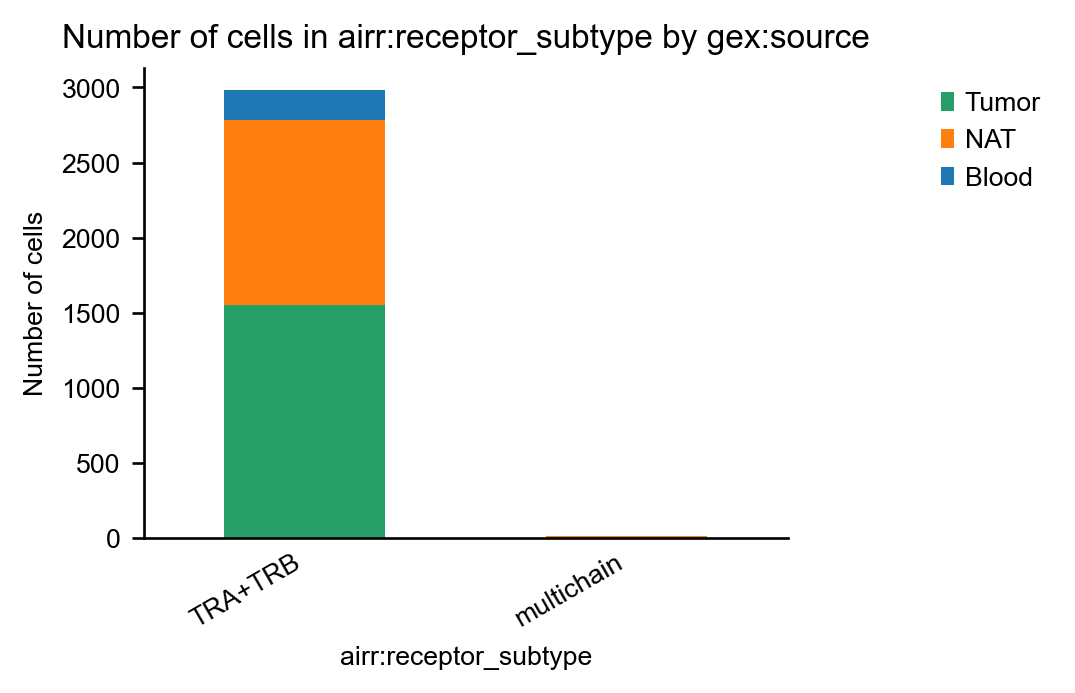
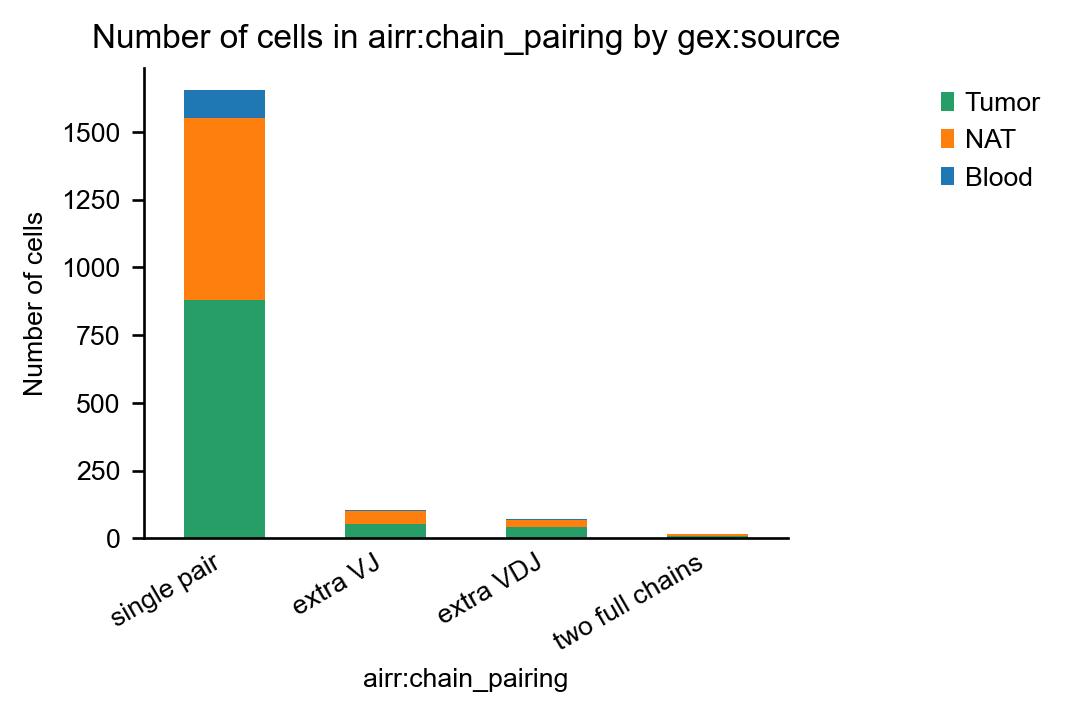
As expected, the dataset contains only α/β T-cell receptors:
Indeed, in this dataset, ~7% of cells have more than one pair of productive T-cell receptors:
print(
"Fraction of cells with more than one pair of TCRs: {:.2f}".format(
np.sum(mdata.obs["airr:chain_pairing"].isin(["extra VJ", "extra VDJ", "two full chains", "multichain"]))
/ mdata["airr"].n_obs
)
)
Fraction of cells with more than one pair of TCRs: 0.07
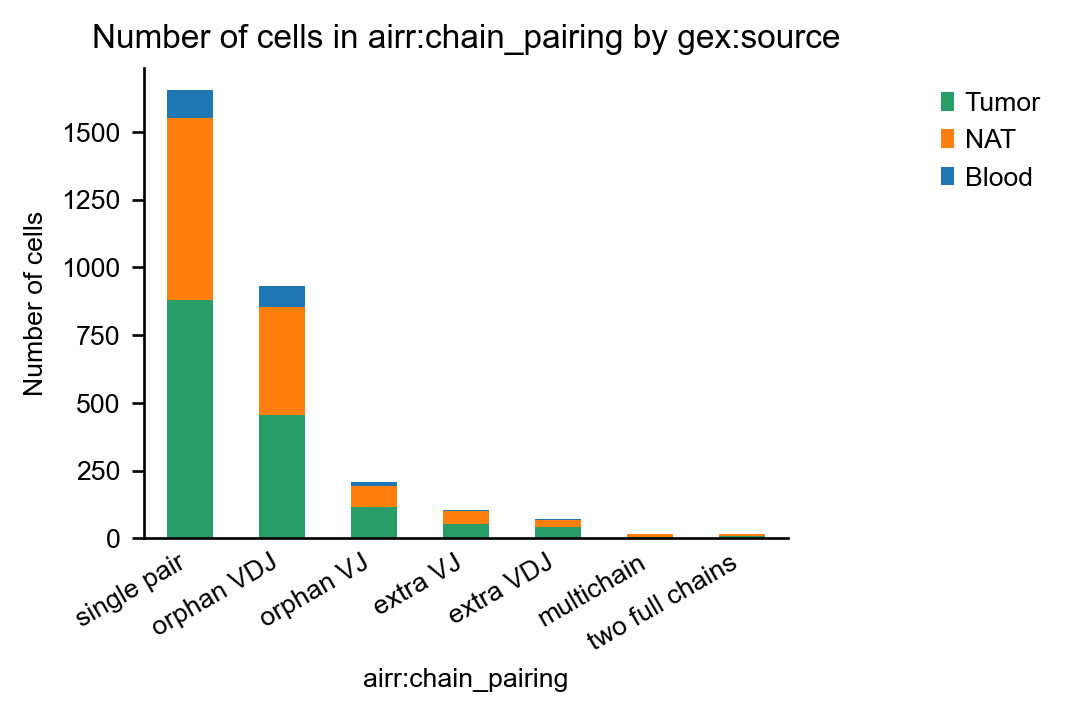
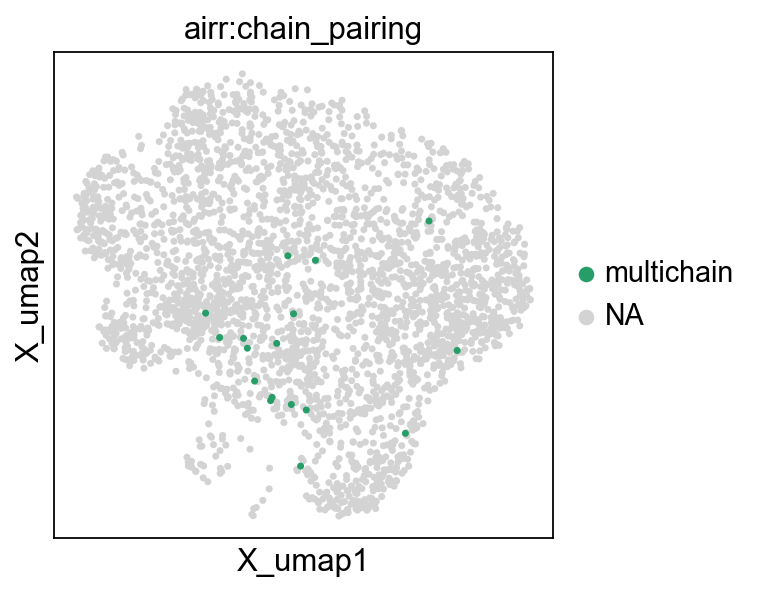
Next, we visualize the Multichain-cells on the UMAP plot and exclude them from downstream analysis:
mu.pp.filter_obs(mdata, "airr:chain_pairing", lambda x: x != "multichain")
Similarly, we can use the chain_pairing information to exclude all cells that don’t have at least
one full pair of receptor sequences:
mu.pp.filter_obs(mdata, "airr:chain_pairing", lambda x: ~np.isin(x, ["orphan VDJ", "orphan VJ"]))
mdata
MuData object with n_obs × n_vars = 1845 × 11850
2 modalities
gex: 1845 x 11850
obs: 'cluster_orig', 'patient', 'sample', 'source', 'n_genes', 'n_counts', 'cluster'
var: 'n_cells', 'highly_variable', 'means', 'dispersions', 'dispersions_norm'
uns: 'cluster_orig_colors', 'log1p', 'hvg', 'pca', 'neighbors', 'gex:sample_colors', 'gex:patient_colors', 'gex:cluster_colors', 'airr:chain_pairing_colors'
obsm: 'X_umap_orig', 'X_pca', 'X_umap'
varm: 'PCs'
obsp: 'distances', 'connectivities'
airr: 1845 x 0
obs: 'high_confidence', 'is_cell', 'clonotype_orig', 'receptor_type', 'receptor_subtype', 'chain_pairing'
uns: 'chain_indices'
obsm: 'airr', 'chain_indices'Finally, we re-create the chain-pairing plot to ensure that the filtering worked as expected:
Define clonotypes and clonotype clusters#
In this section, we will define and visualize clonotypes and clonotype clusters.
Scirpy implements a network-based approach for clonotype definition. The steps to create and visualize the clonotype-network are analogous to the construction of a neighborhood graph from transcriptomics data with Scanpy.
scirpy function |
objective |
|---|---|
Compute sequence-based distance matrices for all VJ and VDJ sequences. |
|
Define clonotypes by nucleotide sequence identity. |
|
Cluster cells by the similarity of their CDR3-sequences. |
|
Compute layout of the clonotype network. |
|
Plot clonotype network colored by different parameters. |
Compute CDR3 neighborhood graph and define clonotypes#
scirpy.pp.ir_dist() computes distances between CDR3 nucleotide (nt) or amino acid (aa) sequences, either based on sequence identity or similarity. It creates two distance matrices: one for all unique VJ sequences and one for all unique VDJ sequences. The distance matrices are added to adata.uns.
The function scirpy.tl.define_clonotypes() matches cells based on the distances of their
VJ and VDJ CDR3-sequences and value of the function parameters dual_ir and receptor_arms. Finally, it
detects connected modules in the graph and annotates them as clonotypes. This will add a clone_id and
clone_id_size column to adata.obs.
The dual_ir parameter defines how scirpy handles cells with more than one pair of receptors. The default value is any which implies that cells with any of their primary or secondary receptor chain matching will be considered to be of the same clonotype.
Here, we define clonotypes based on nt-sequence identity. In a later step, we will define clonotype clusters based on amino-acid similarity.
# using default parameters, `ir_dist` will compute nucleotide sequence identity
ir.pp.ir_dist(mdata)
ir.tl.define_clonotypes(mdata, receptor_arms="all", dual_ir="primary_only")
Computing sequence x sequence distance matrix for VJ sequences.
Computing sequence x sequence distance matrix for VDJ sequences.
Initializing lookup tables.
Computing clonotype x clonotype distances.
Stored result in `mdata.obs["airr:clone_id"]`.
Stored result in `mdata.obs["airr:clone_id_size"]`.
To visualize the network we first call scirpy.tl.clonotype_network() to compute the layout.
We can then visualize it using scirpy.pl.clonotype_network(). We recommend setting the
min_cells parameter to >=2, to prevent the singleton clonotypes from cluttering the network.
ir.tl.clonotype_network(mdata, min_cells=2)
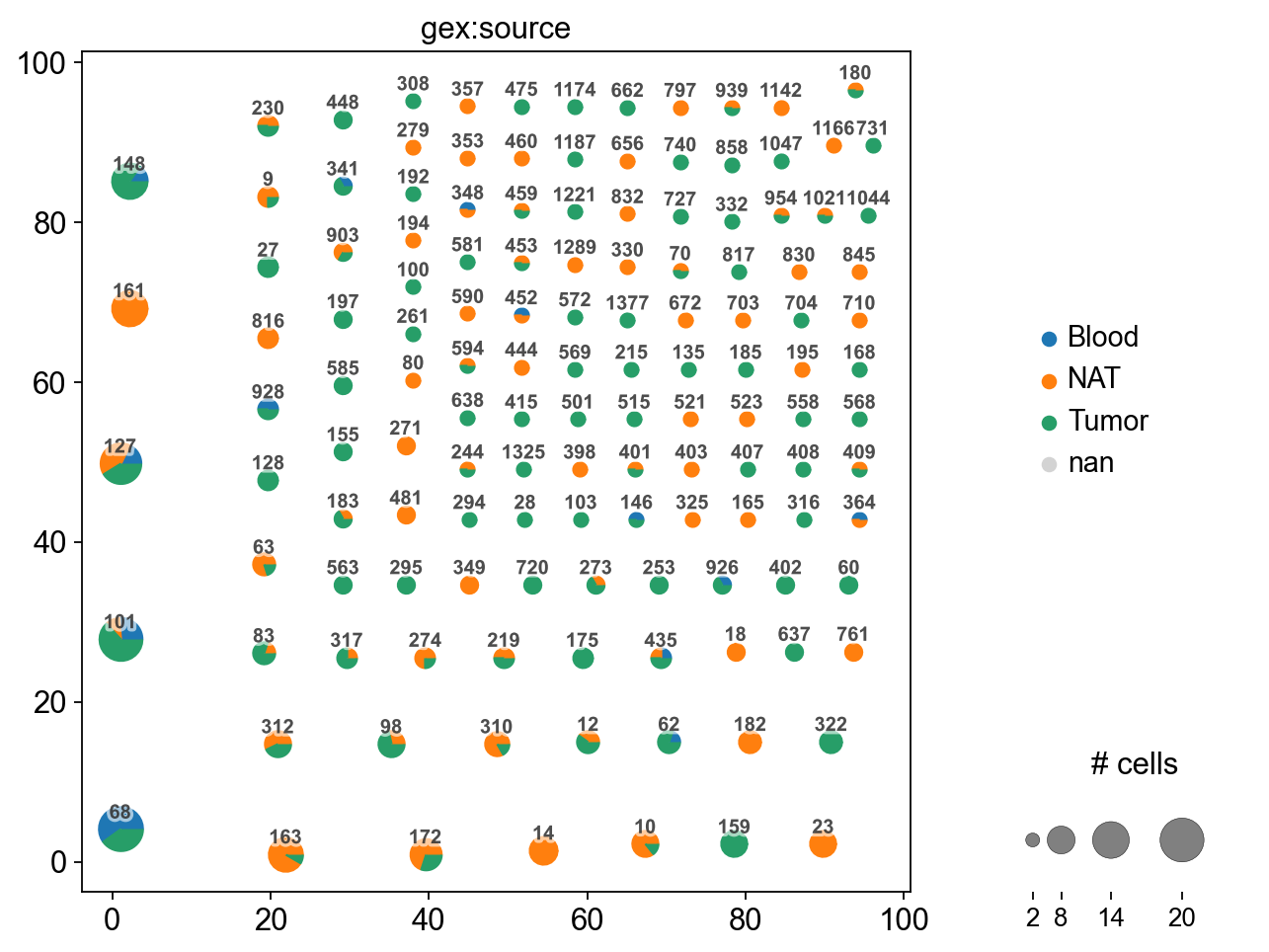
The resulting plot is a network, where each dot represents cells with identical receptor configurations. As we define clonotypes as cells with identical CDR3-sequences, each dot is also a clonotype. For each clonotype, the numeric clonotype id is shown in the graph. The size of each dot refers to the number of cells with the same receptor configurations. Categorical variables can be visualized as pie charts.
mdata.obs.groupby("gex:source", dropna=False).size()
gex:source
Blood 107
NAT 756
Tumor 982
dtype: int64
Re-compute CDR3 neighborhood graph and define clonotype clusters#
We can now re-compute the clonotype network based on amino-acid sequence similarity and define clonotype clusters.
To this end, we need to change set metric="alignment" and specify a cutoff parameter.
The distance is based on the BLOSUM62 matrix.
For instance, a distance of 10 is equivalent to 2 Rs mutating into N.
This appoach was initially proposed as TCRdist by Dash et al. [DFGH+17].
Speeding up TCR distance calculation
Scirpy provides alternative distance metrics that are similar to "alignment", but a lot faster:
metric="tcrdist"is an implementation of tcrdist3 within scirpy. The scores are calculated differently, but it gives very similar results compared tometric="alignment". See alsoscirpy.ir_dist.metrics.TCRdistDistanceCalculator.metric="fastalignment"uses a heuristic to speed up the"alignment"metric at the cost of a few false-negatives (i.e. sequence pairs that are actually below the distance cutoff, but are removed during a pre-filtering step). With default parameters, the false-negative rate (of all sequence pairs actually below the cutoff) was ~2% on thescirpy.datasets.wu2020()dataset. See alsoscirpy.ir_dist.metrics.FastAlignmentDistanceCalculator.
All cells with a distance between their CDR3 sequences lower than cutoff will be connected in the network.
ir.pp.ir_dist(
mdata,
metric="alignment",
sequence="aa",
cutoff=15,
)
Computing sequence x sequence distance matrix for VJ sequences.
Computing sequence x sequence distance matrix for VDJ sequences.
ir.tl.define_clonotype_clusters(mdata, sequence="aa", metric="alignment", receptor_arms="all", dual_ir="any")
Initializing lookup tables.
Computing clonotype x clonotype distances.
Stored result in `mdata.obs["airr:cc_aa_alignment"]`.
Stored result in `mdata.obs["airr:cc_aa_alignment_size"]`.
ir.tl.clonotype_network(mdata, min_cells=3, sequence="aa", metric="alignment")
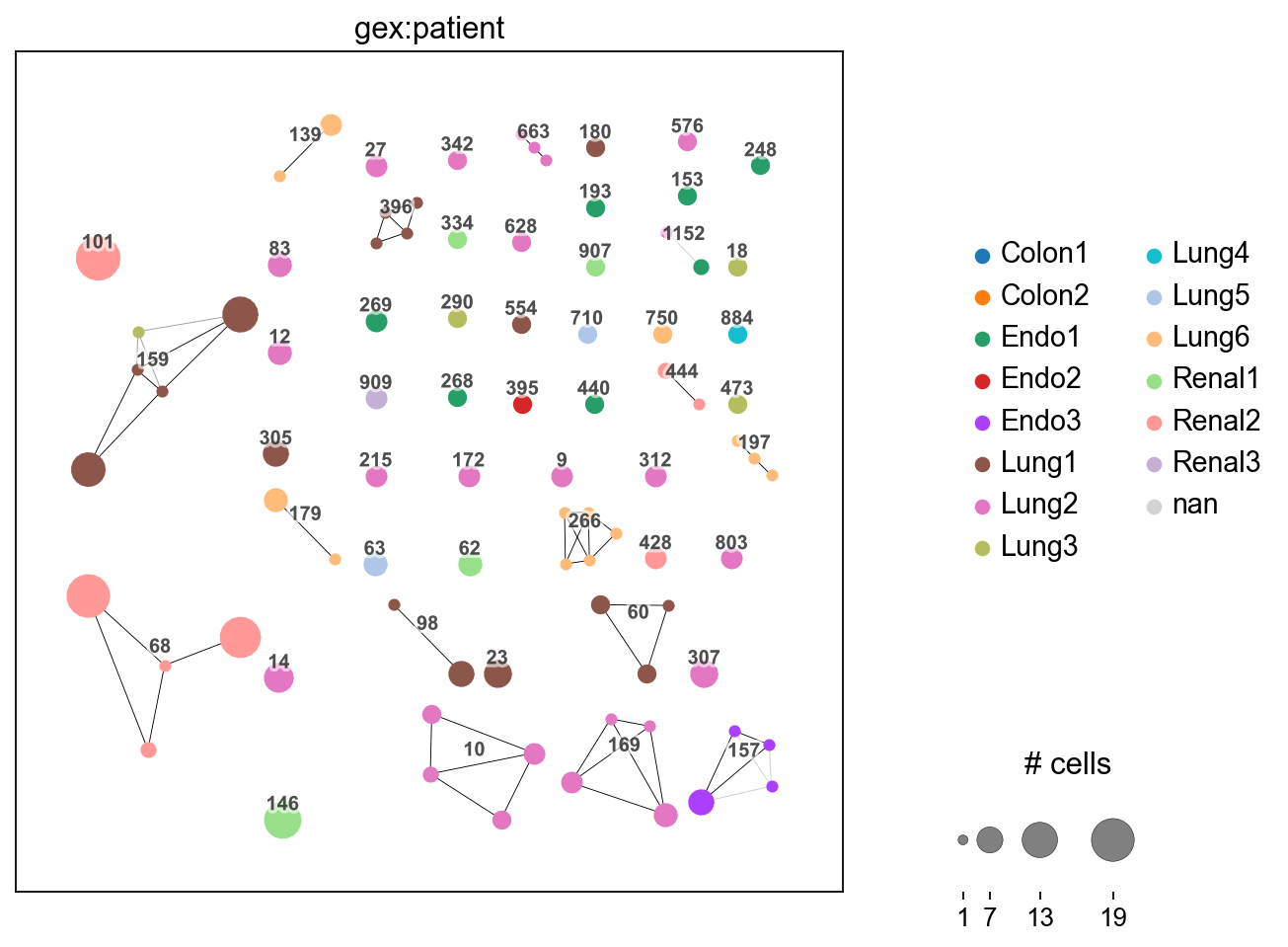
Compared to the previous plot, we observere several connected dots. Each fully connected subnetwork represents a “clonotype cluster”, each dot still represents cells with identical receptor configurations.
The dots are colored by patient. We observe, that for instance, clonotypes 101 and 68 (left top and bottom) are private, i.e. they contain cells from a single patient only. On the other hand, clonotype 159 (left middle) is
public, i.e. it is shared across patients Lung1 and Lung3.
_ = ir.pl.clonotype_network(mdata, color="gex:patient", label_fontsize=9, panel_size=(7, 7), base_size=20)
Note
Since v0.13, AIRR data is not by default included in obs. We can retrieve a per-cell data frame with AIRR information
using scirpy.get.airr(), or temporarily add columns to obs using the scirpy.get.airr_context() context
manager. For more information, see Accessing AIRR data.
We can now extract information (e.g. CDR3-sequences) from a specific clonotype cluster by subsetting MuData.
By extracting the CDR3 sequences of clonotype cluster 159, we retreive five different receptor configurations with different numbers of cells, corresponding to the five points in the graph.
with ir.get.airr_context(mdata, "junction_aa", ["VJ_1", "VDJ_1", "VJ_2", "VDJ_2"]):
cdr3_ct_159 = (
# TODO astype(str) is required due to a bug in pandas ignoring `dropna=False`. It seems fixed in pandas 2.x
mdata.obs.loc[lambda x: x["airr:cc_aa_alignment"] == "159"]
.astype(str)
.groupby(
[
"VJ_1_junction_aa",
"VDJ_1_junction_aa",
"VJ_2_junction_aa",
"VDJ_2_junction_aa",
"airr:receptor_subtype",
],
observed=True,
dropna=False,
)
.size()
.reset_index(name="n_cells")
)
cdr3_ct_159
| VJ_1_junction_aa | VDJ_1_junction_aa | VJ_2_junction_aa | VDJ_2_junction_aa | airr:receptor_subtype | n_cells | |
|---|---|---|---|---|---|---|
| 0 | CAGKAGNTGKLIF | CASSYQGSTEAFF | nan | nan | TRA+TRB | 1 |
| 1 | CAGKSGNTGKLIF | CASSYQGATEAFF | CATDPRRSTGNQFYF | nan | TRA+TRB | 1 |
| 2 | CAGKSGNTGKLIF | CASSYQGATEAFF | nan | nan | TRA+TRB | 12 |
| 3 | CATDPRRSTGNQFYF | CASSYQGATEAFF | CAGKSGNTGKLIF | nan | TRA+TRB | 1 |
| 4 | CATDPRRSTGNQFYF | CASSYQGATEAFF | nan | nan | TRA+TRB | 11 |
Including the V-gene in clonotype definition#
Using the paramter use_v_gene in define_clonotypes(), we can enforce
clonotypes (or clonotype clusters) to have the same V-gene, and, therefore, the same CDR1 and 2
regions. Let’s look for clonotype clusters with different V genes:
ir.tl.define_clonotype_clusters(
mdata,
sequence="aa",
metric="alignment",
receptor_arms="all",
dual_ir="any",
same_v_gene=True,
key_added="cc_aa_alignment_same_v",
)
Initializing lookup tables.
Computing clonotype x clonotype distances.
Stored result in `mdata.obs["airr:cc_aa_alignment_same_v"]`.
Stored result in `mdata.obs["airr:cc_aa_alignment_same_v_size"]`.
# find clonotypes with more than one `clonotype_same_v`
ct_different_v = mdata.obs.groupby("airr:cc_aa_alignment").apply(
lambda x: x["airr:cc_aa_alignment_same_v"].nunique() > 1
)
ct_different_v = ct_different_v[ct_different_v].index.values.tolist()
ct_different_v
['280', '765']
Here, we see that the clonotype clusters 280 and 765 get split into (280, 788) and (765, 1071), respectively, when the same_v_gene flag is set.
with ir.get.airr_context(mdata, "v_call", ["VJ_1", "VDJ_1"]):
ct_different_v_df = (
mdata.obs.loc[
lambda x: x["airr:cc_aa_alignment"].isin(ct_different_v),
[
"airr:cc_aa_alignment",
"airr:cc_aa_alignment_same_v",
"VJ_1_v_call",
"VDJ_1_v_call",
],
]
.sort_values("airr:cc_aa_alignment")
.drop_duplicates()
.reset_index(drop=True)
)
ct_different_v_df
| airr:cc_aa_alignment | airr:cc_aa_alignment_same_v | VJ_1_v_call | VDJ_1_v_call | |
|---|---|---|---|---|
| 0 | 280 | 280 | TRAV8-6 | TRBV6-6 |
| 1 | 280 | 788 | TRAV8-3 | TRBV9 |
| 2 | 765 | 765 | TRAV21 | TRBV6-6 |
| 3 | 765 | 1071 | TRAV21 | TRBV6-5 |
Clonotype analysis#
Clonal expansion#
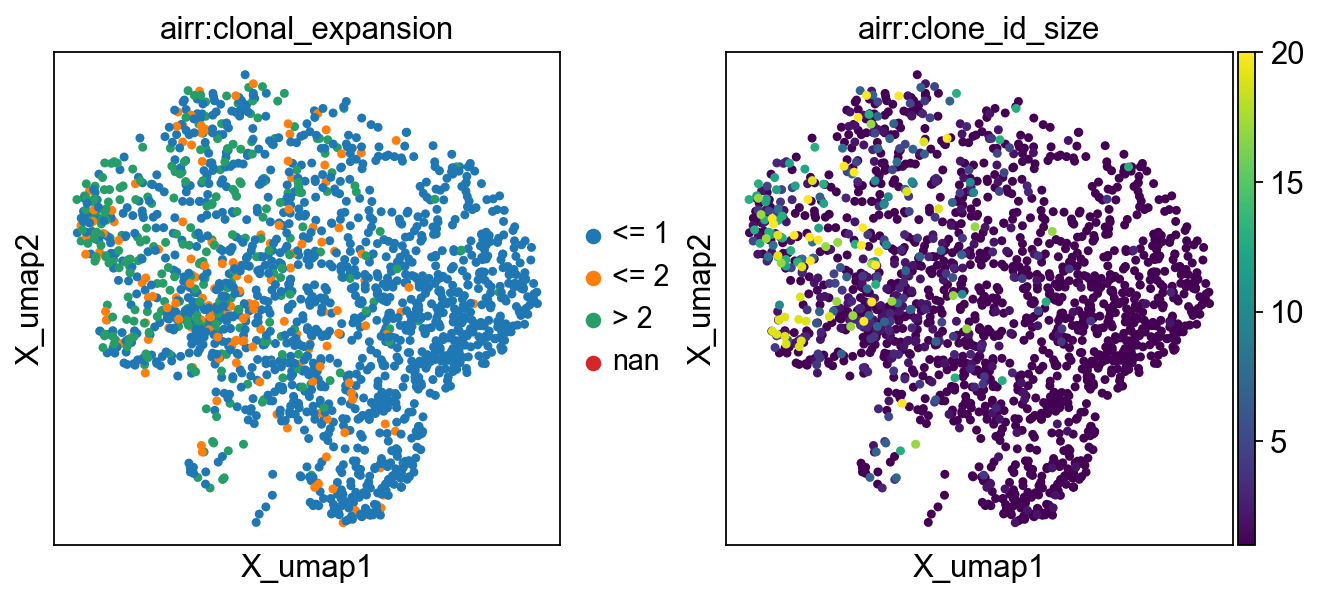
Let’s visualize the number of expanded clonotypes (i.e. clonotypes consisting
of more than one cell) by cell-type. The first option is to add a column with the scirpy.tl.clonal_expansion()
to adata.obs and overlay it on the UMAP plot.
ir.tl.clonal_expansion(mdata)
Stored result in `mdata.obs["airr:clonal_expansion"]`.
clonal_expansion refers to expansion categories, i.e singleton clonotypes, clonotypes with 2 cells and more than 2 cells.
The clonotype_size refers to the absolute number of cells in a clonotype.
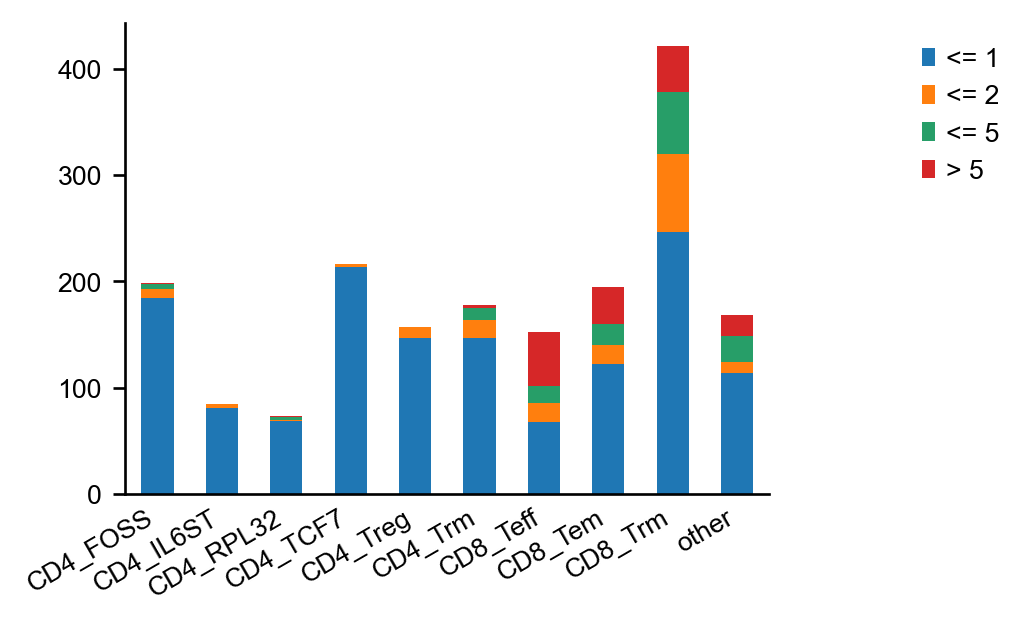
The second option is to show the number of cells belonging to an expanded clonotype per category
in a stacked bar plot, using the scirpy.pl.clonal_expansion() plotting function.
_ = ir.pl.clonal_expansion(mdata, target_col="clone_id", groupby="gex:cluster", breakpoints=(1, 2, 5), normalize=False)
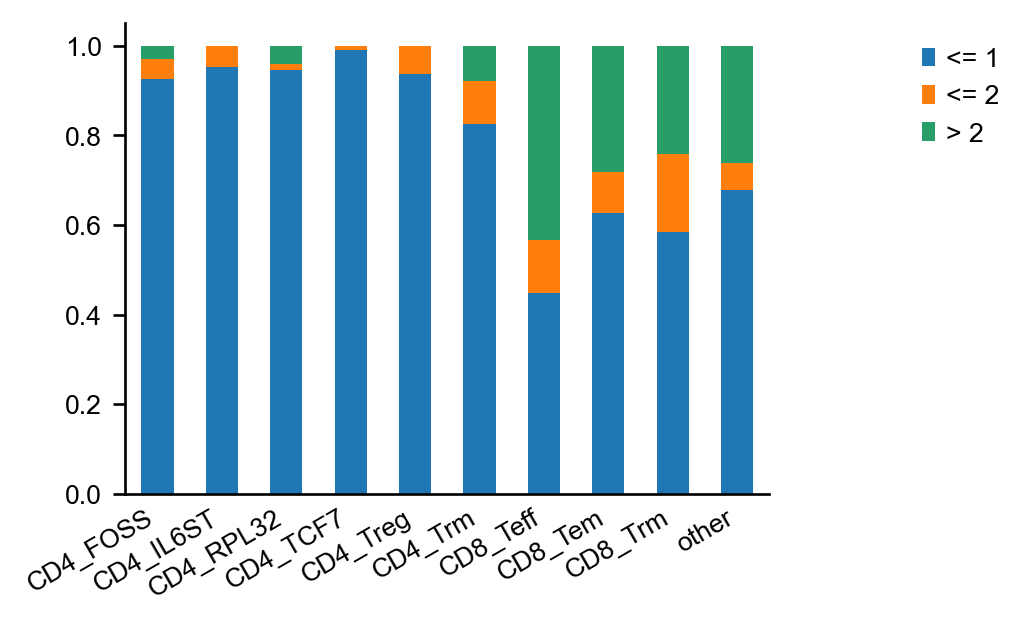
The same plot, normalized to cluster size. Clonal expansion is a sign of positive selection for a certain, reactive T-cell clone. It, therefore, makes sense that CD8+ effector T-cells have the largest fraction of expanded clonotypes.
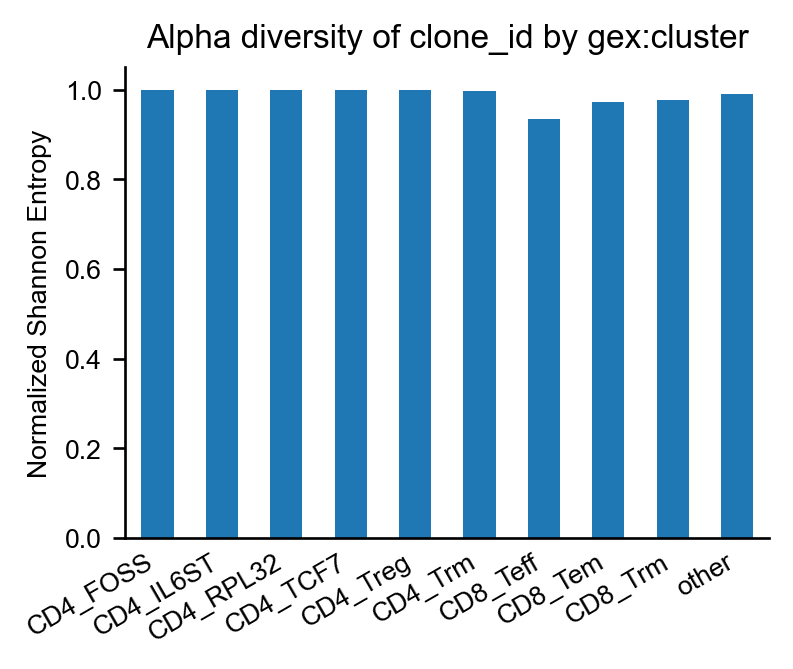
Expectedly, the CD8+ effector T cells have the largest fraction of expanded clonotypes.
Consistent with this observation, they have the lowest scirpy.pl.alpha_diversity() of clonotypes.
Clonotype abundance#
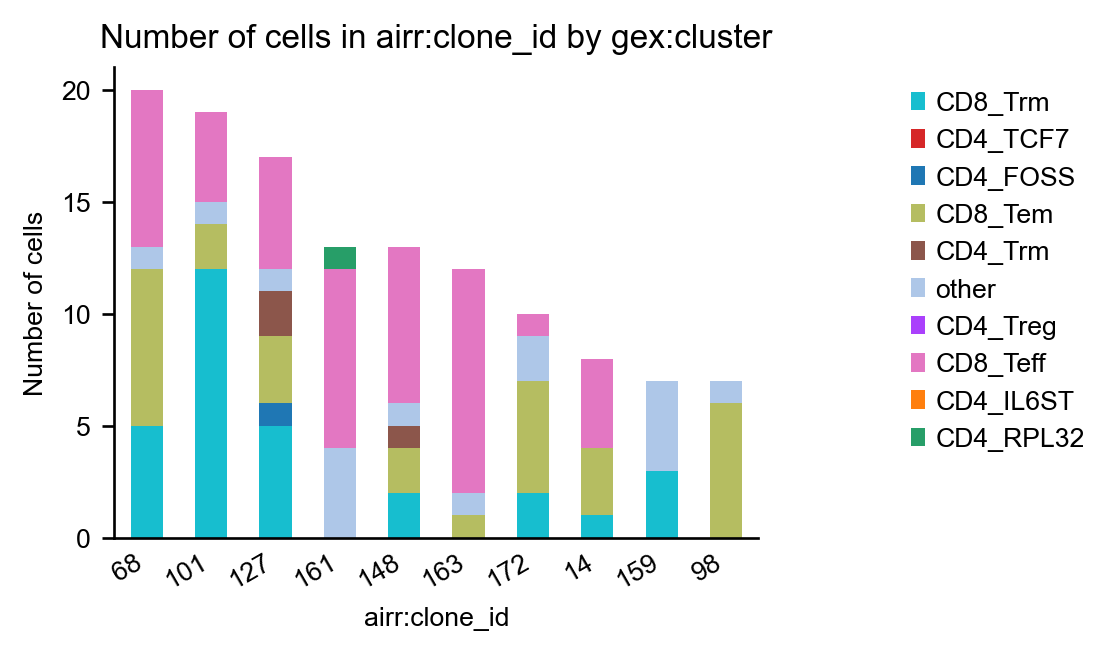
The function scirpy.pl.group_abundance() allows us to create bar charts for
arbitrary categorial from obs. Here, we use it to show the distribution of them
ten largest clonotypes across the cell-type clusters.
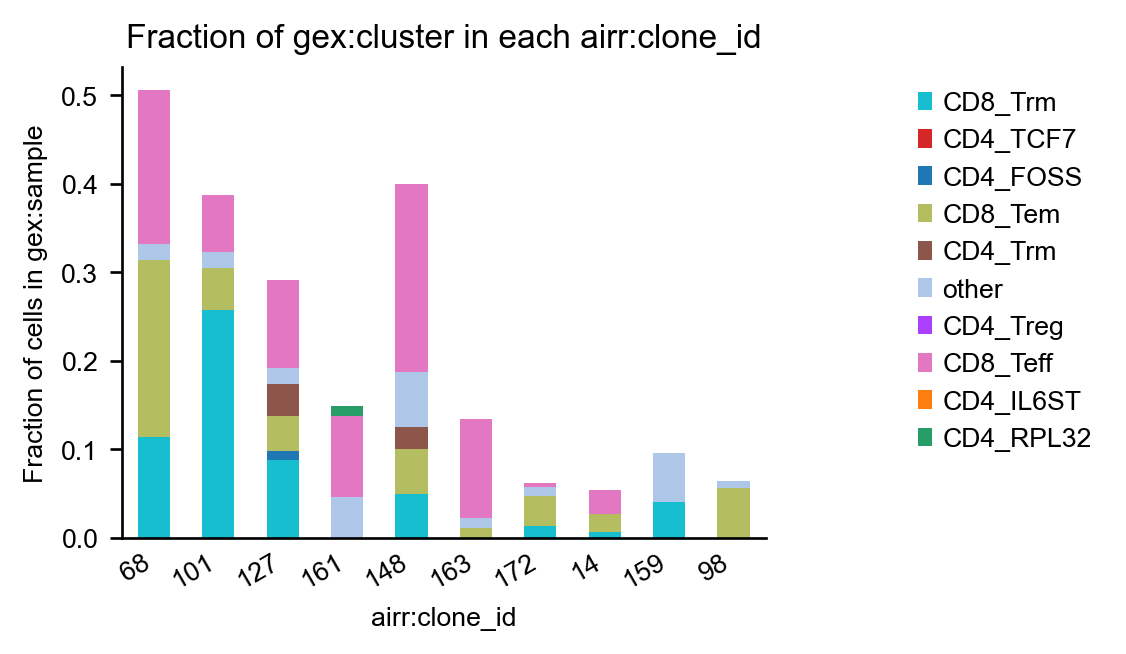
It might be beneficial to normalize the counts to the number of cells per sample to mitigate biases due to different sample sizes:
_ = ir.pl.group_abundance(
mdata,
groupby="airr:clone_id",
target_col="gex:cluster",
max_cols=10,
normalize="gex:sample",
)
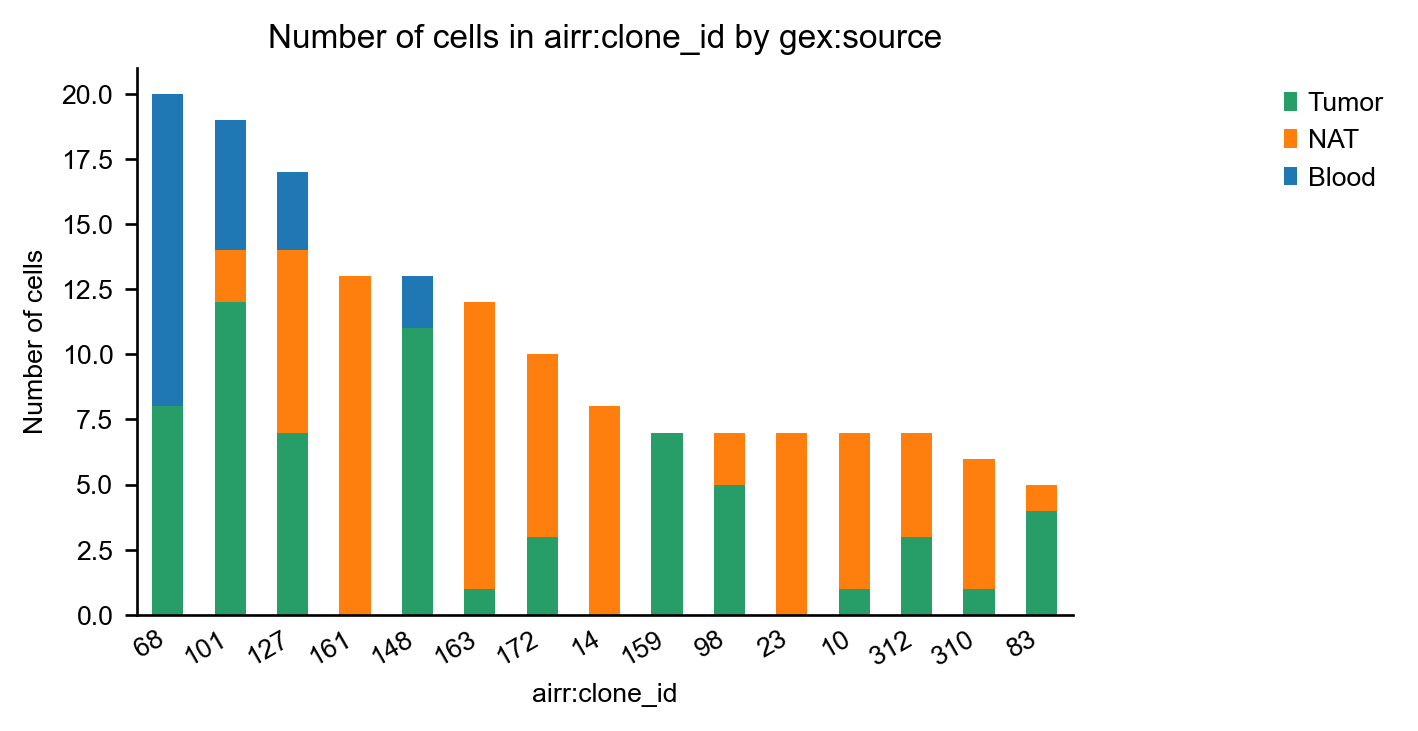
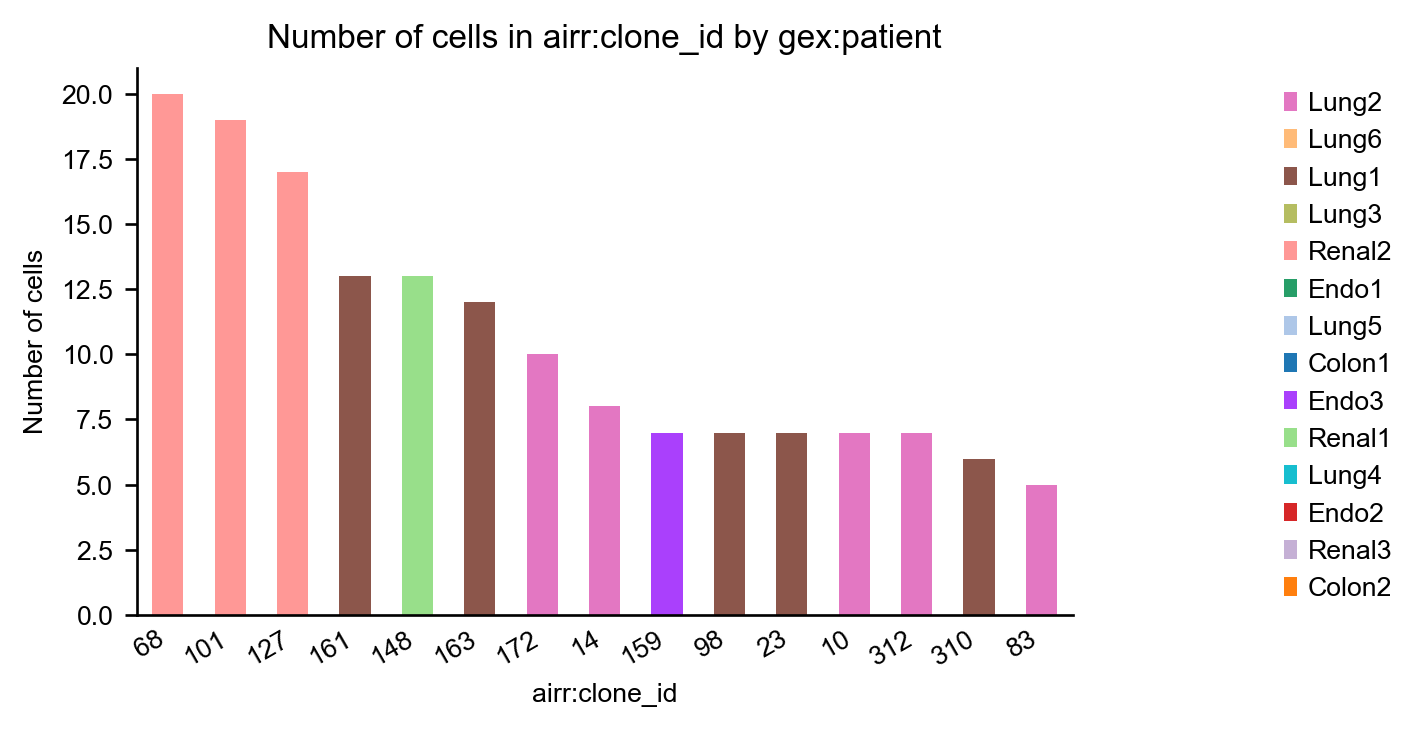
Coloring the bars by patient gives us information about public and private clonotypes: Some clonotypes are private, i.e. specific to a certain tissue, others are public, i.e. they are shared across different tissues.
_ = ir.pl.group_abundance(mdata, groupby="airr:clone_id", target_col="gex:source", max_cols=15, figsize=(5, 3))
However, clonotypes that are shared between patients are rare:
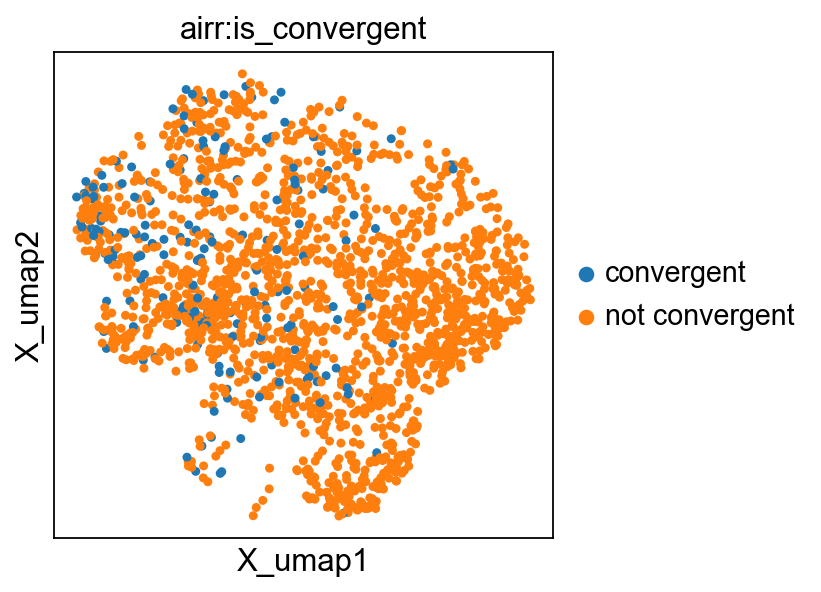
Convergent evolution#
By comparing two levels of clonotype definitions (e.g. based on nucleotide sequences and based on amino-acid sequences), we can identify receptors that are subject to convergent evolution. By that, we mean receptors that (likely) recognize the same antigen but have evolved from different clones.
ir.tl.clonotype_convergence(mdata, key_coarse="cc_aa_alignment", key_fine="clone_id")
Stored result in `mdata.obs["airr:is_convergent"]`.
Gene usage#
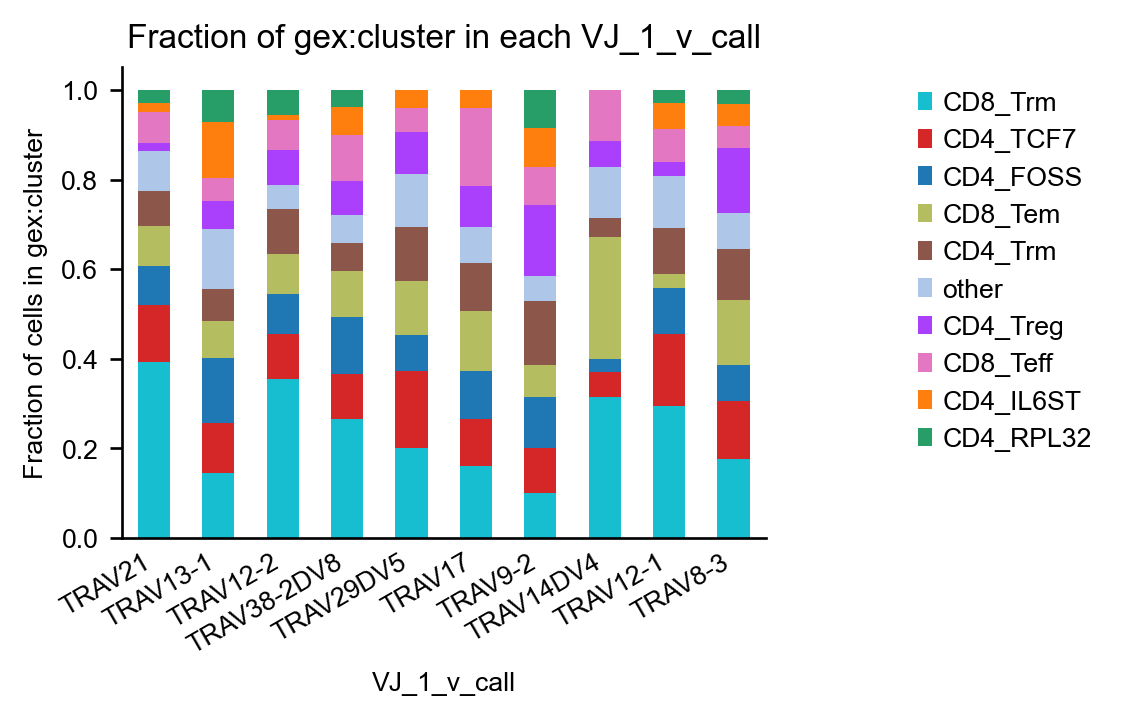
scirpy.tl.group_abundance() can also give us some information on VDJ usage.
We first add information about v_call to obs using airr_context() and then generate a bar plot by the VJ_1_v_call variable.
We use max_col to limit the plot to the 10 most abundant V-genes.
with ir.get.airr_context(mdata, "v_call"):
ir.pl.group_abundance(
mdata,
groupby="VJ_1_v_call",
target_col="gex:cluster",
normalize=True,
max_cols=10,
)
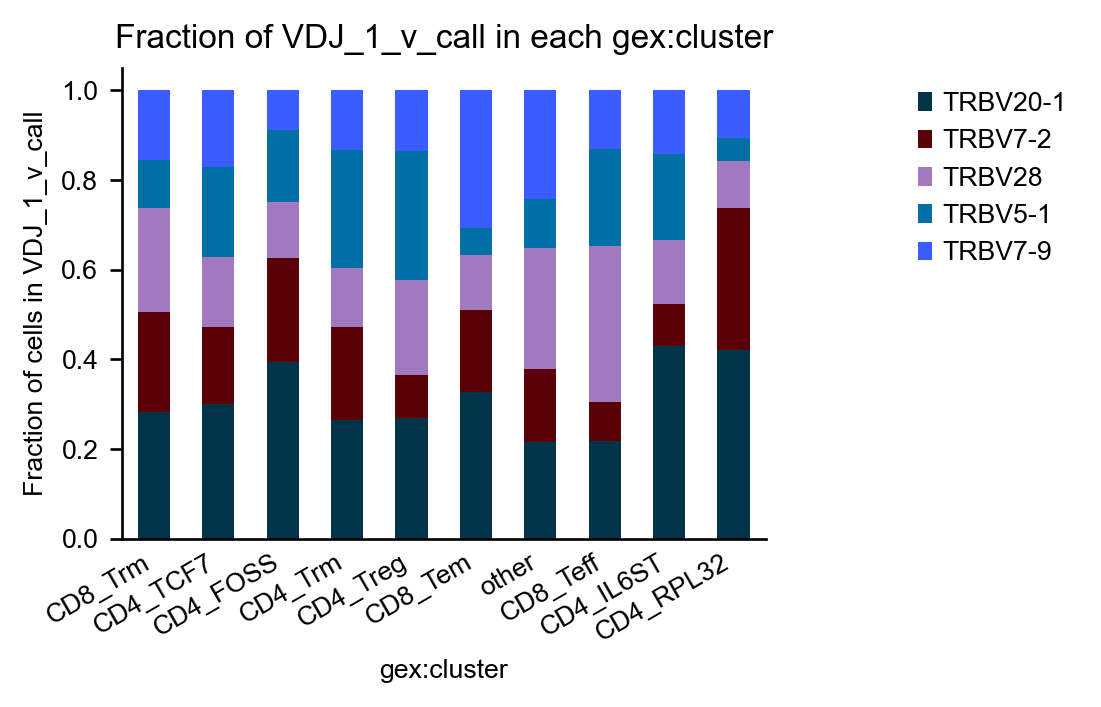
We can pre-select groups by filtering adata:
with ir.get.airr_context(mdata, "v_call"):
ir.pl.group_abundance(
mdata[
mdata.obs["VDJ_1_v_call"].isin(["TRBV20-1", "TRBV7-2", "TRBV28", "TRBV5-1", "TRBV7-9"]),
:,
],
groupby="gex:cluster",
target_col="VDJ_1_v_call",
normalize=True,
)
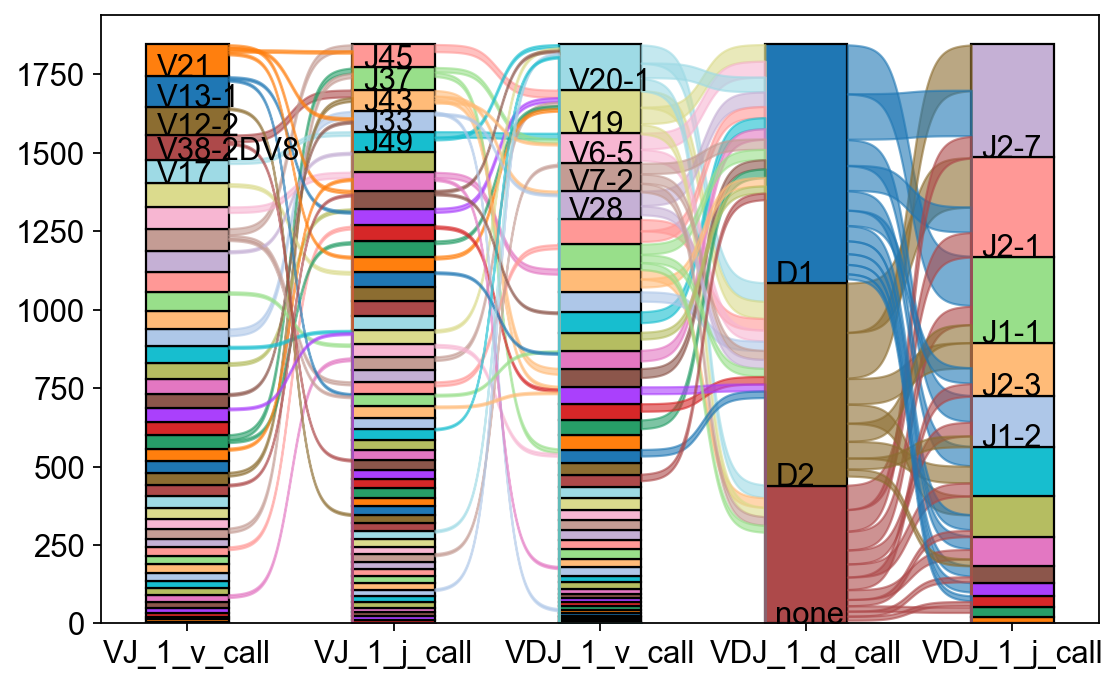
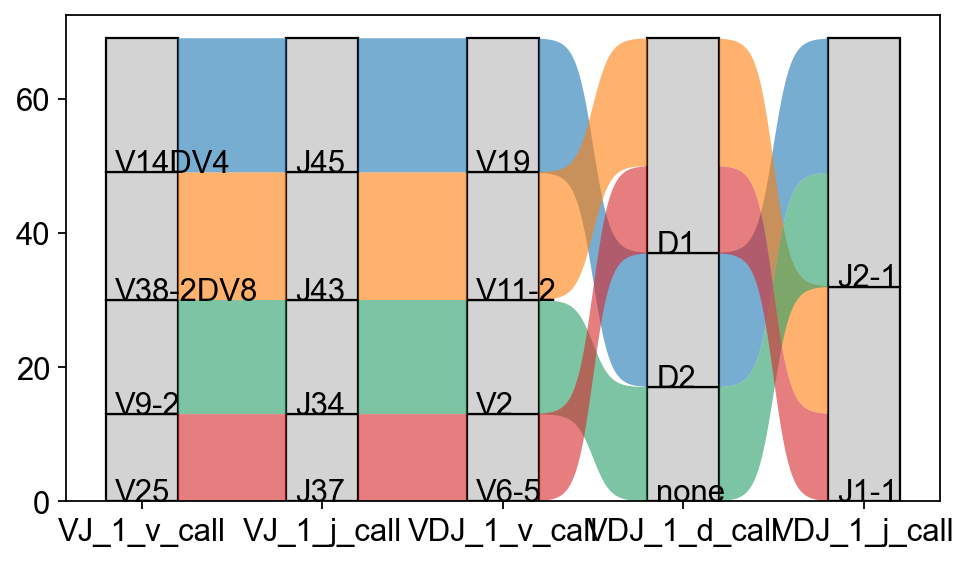
The exact combinations of VDJ genes can be visualized as a Sankey-plot using scirpy.pl.vdj_usage().
_ = ir.pl.vdj_usage(
mdata,
full_combination=False,
max_segments=None,
max_ribbons=30,
fig_kws={"figsize": (8, 5)},
)
We can also use this plot to investigate the exact VDJ composition of one (or several) clonotypes:
ir.pl.vdj_usage(
mdata[mdata.obs["airr:clone_id"].isin(["68", "101", "127", "161"]), :],
max_ribbons=None,
max_segments=100,
)
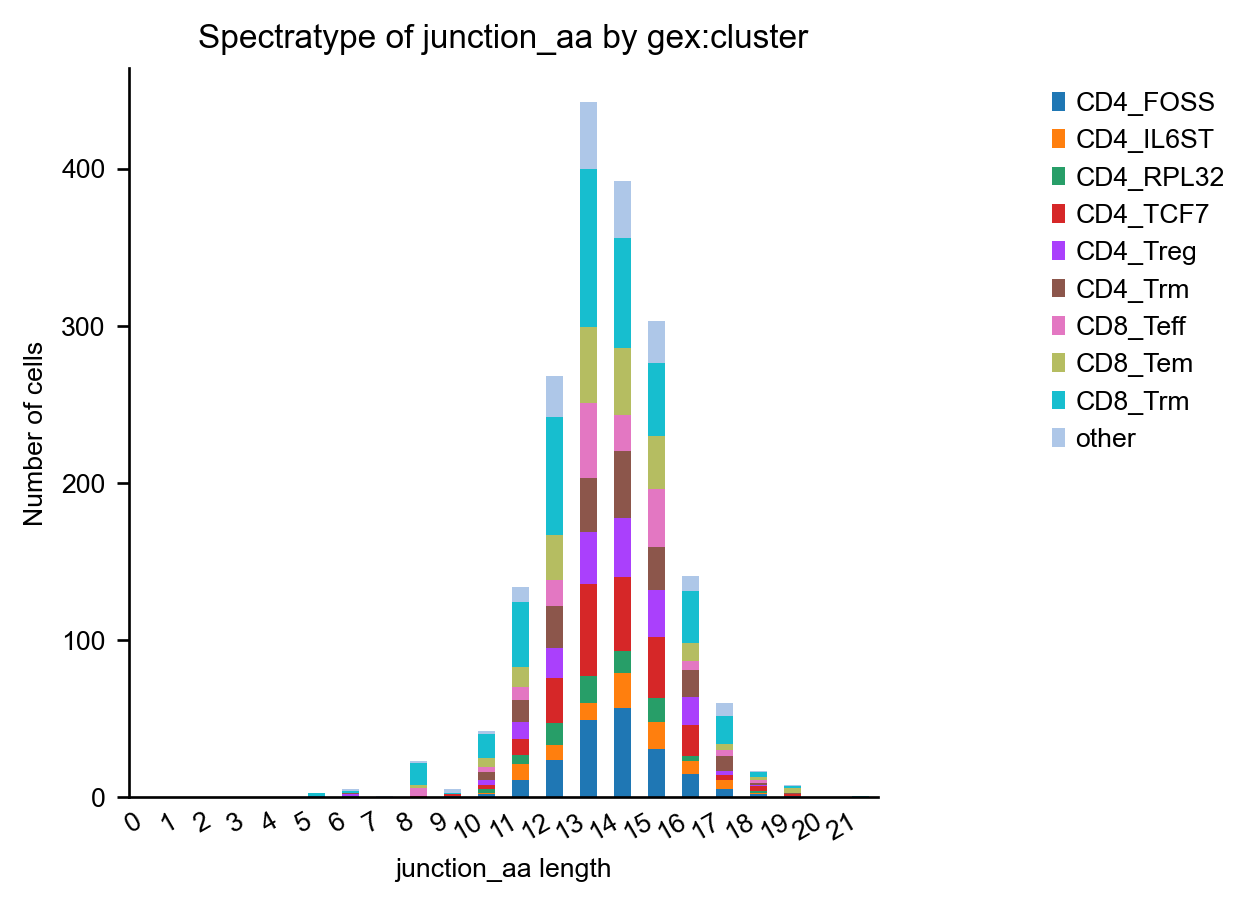
Spectratype plots#
spectratype() plots give us information about the length distribution of CDR3 regions.
ir.pl.spectratype(mdata, color="gex:cluster", viztype="bar", fig_kws={"dpi": 120})
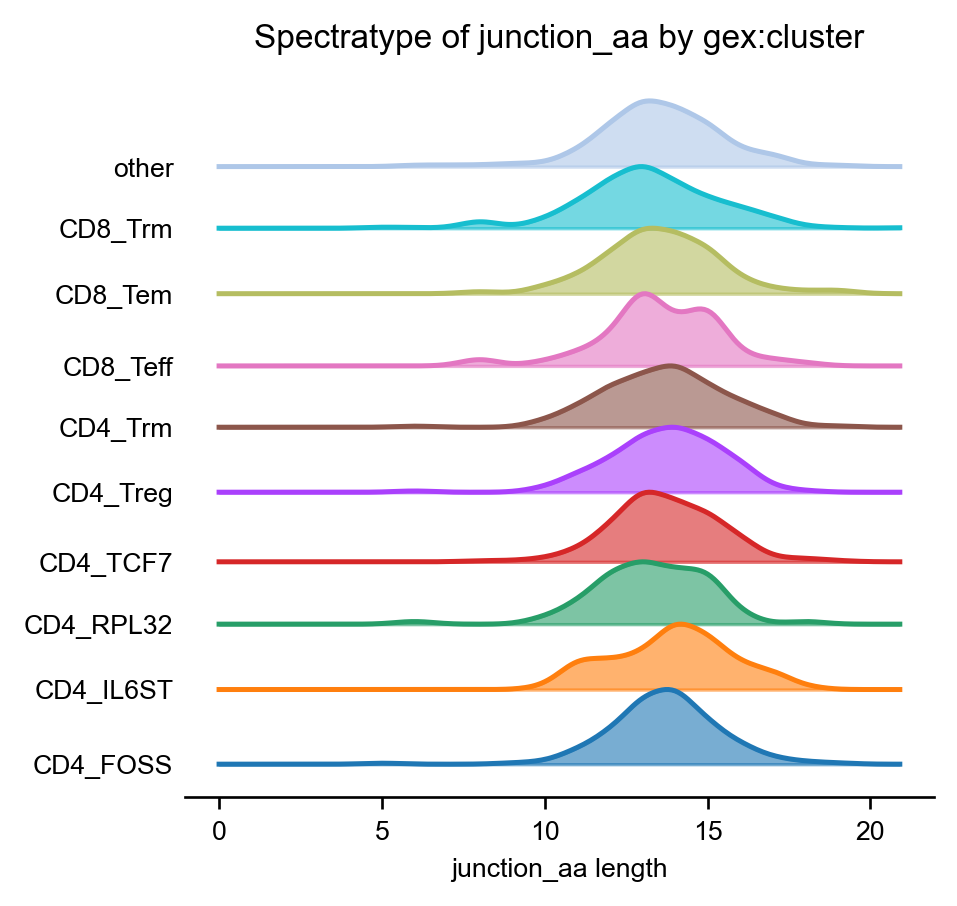
The same chart visualized as “ridge”-plot:
ir.pl.spectratype(
mdata,
color="gex:cluster",
viztype="curve",
curve_layout="shifted",
fig_kws={"dpi": 120},
kde_kws={"kde_norm": False},
)
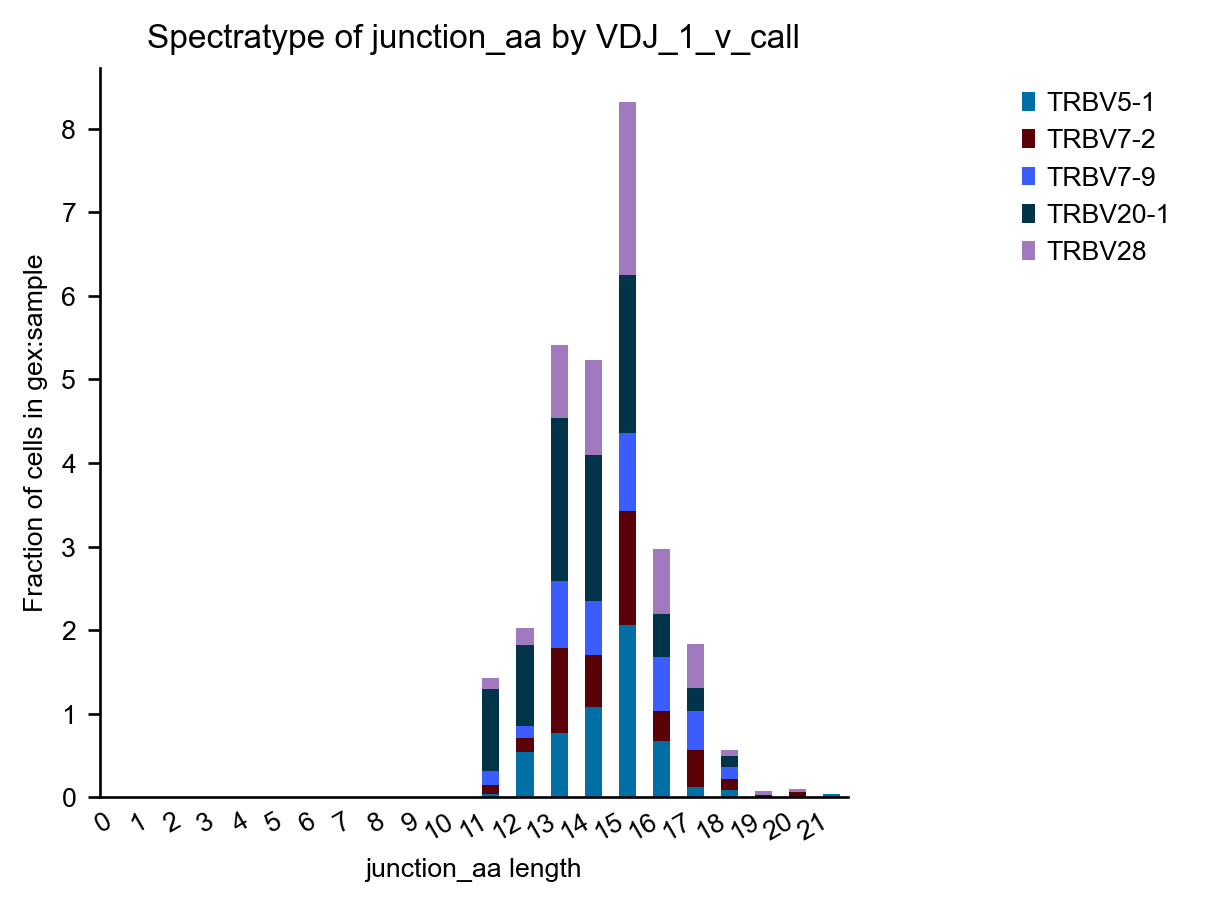
A spectratype-plot by gene usage. To pre-select specific genes, we can simply filter the mdata object before plotting.
Comparing repertoires#
Repertoire simlarity and overlaps#
Overlaps in the adaptive immune receptor repertoire of samples or sample groups enables to pinpoint important clonotype groups, as well as to provide a measure of similarity between samples.
Running Scirpy’s repertoire_overlap() tool creates a matrix featuring the abundance of clonotypes in each sample. Additionally, it also computes a (Jaccard) distance matrix of samples as well as the linkage of hierarchical clustering.
df, dst, lk = ir.tl.repertoire_overlap(mdata, "gex:sample", inplace=False)
df.head()
| clone_id | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | ... | 1516 | 1517 | 1518 | 1519 | 1520 | 1521 | 1522 | 1523 | 1524 | 1525 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| gex:sample | |||||||||||||||||||||
| RN2 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| LN1 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| LN2 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 3.0 | ... | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 |
| LN4 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| LT6 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | ... | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 |
5 rows × 1526 columns
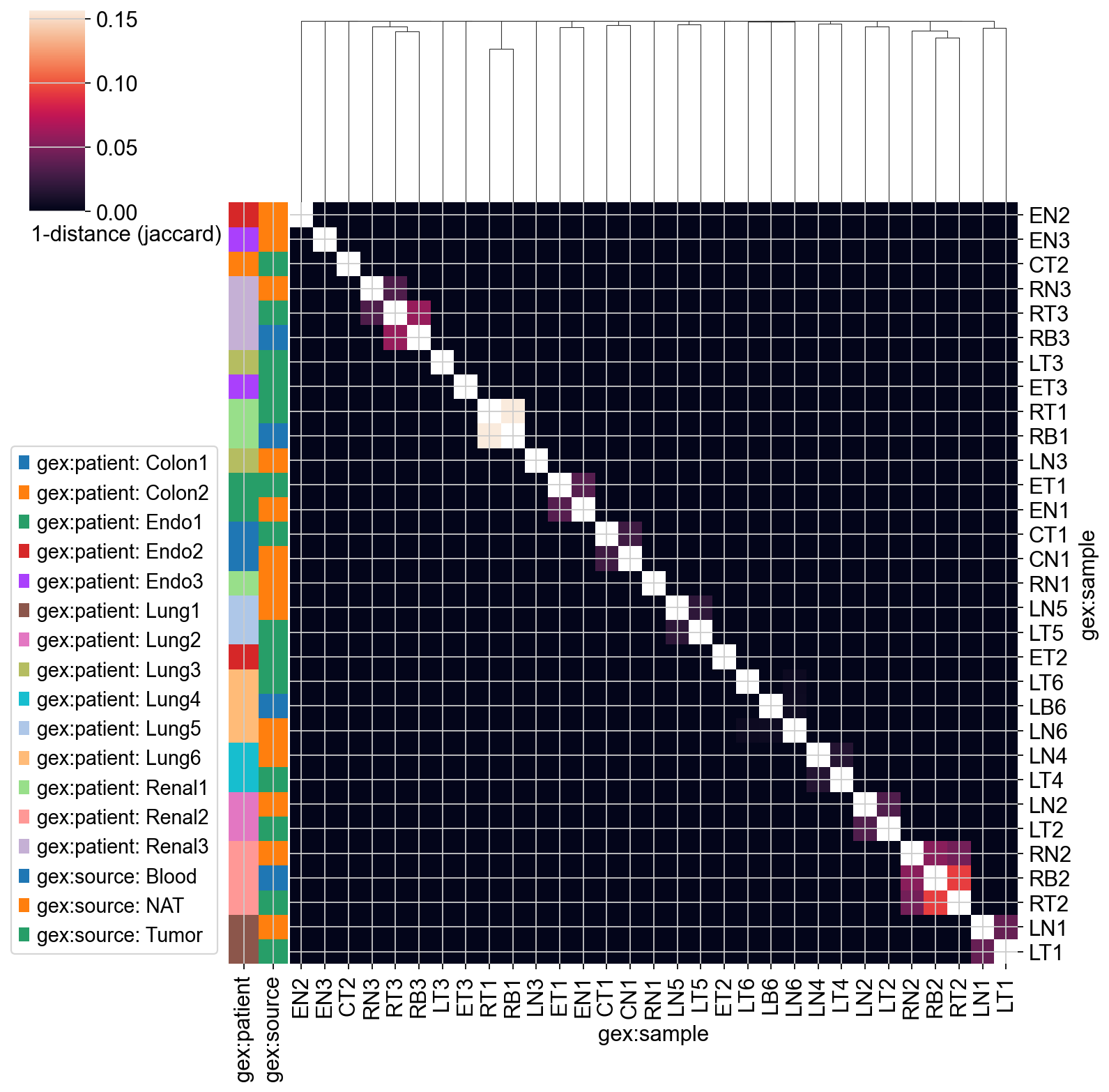
The distance matrix can be shown as a heatmap, while samples are reordered based on hierarchical clustering.
_ = ir.pl.repertoire_overlap(
mdata,
"gex:sample",
heatmap_cats=["gex:patient", "gex:source"],
yticklabels=True,
xticklabels=True,
)
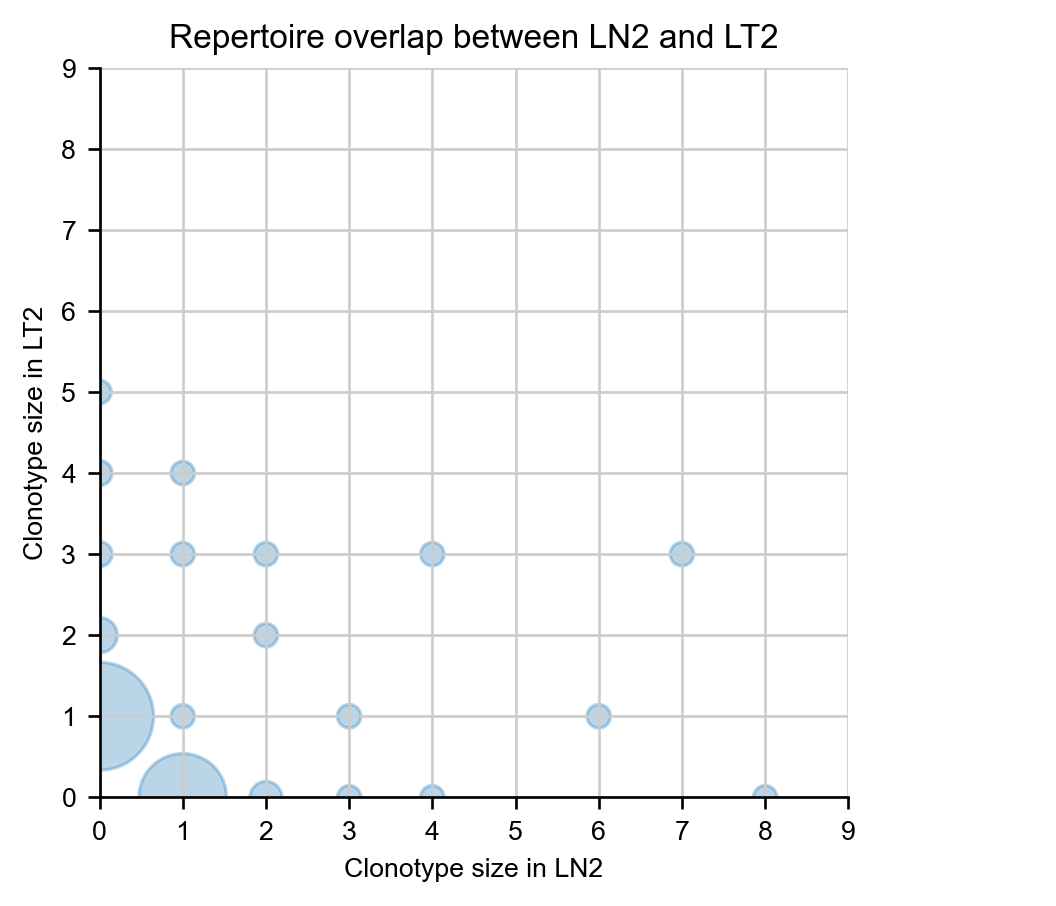
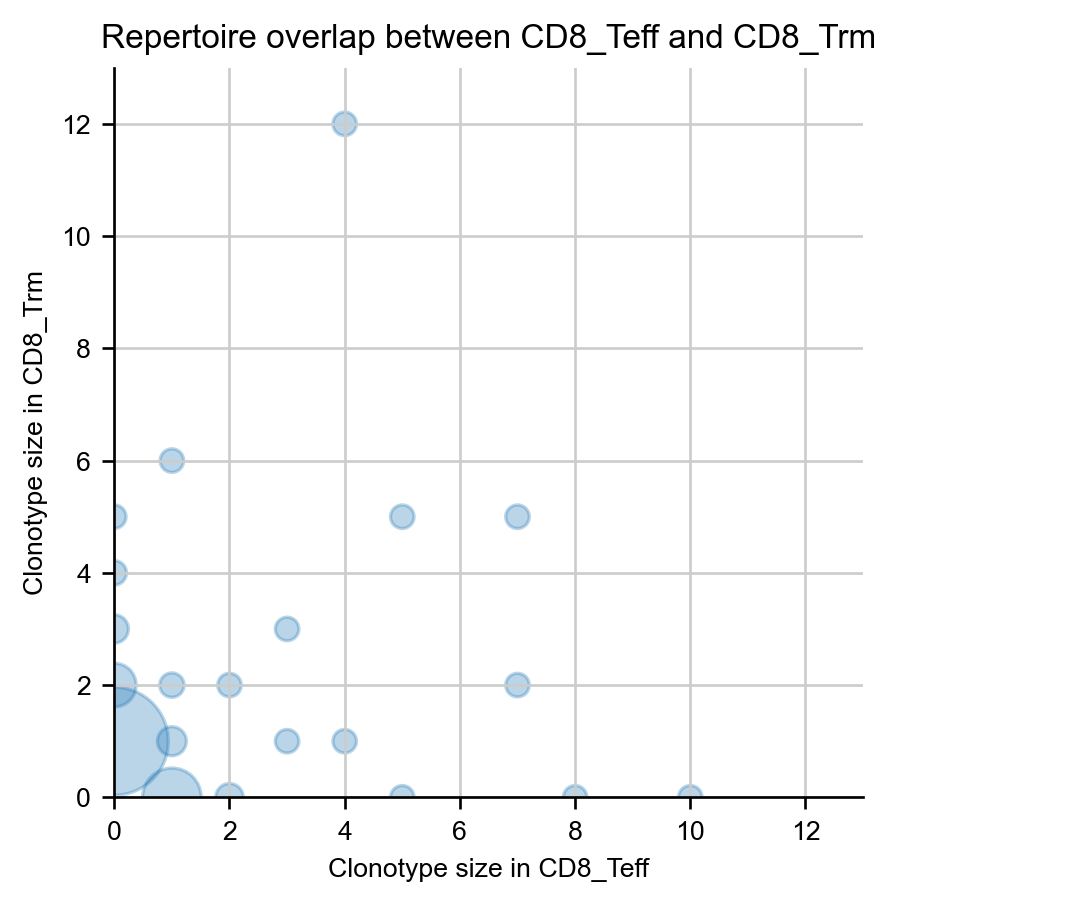
A specific pair of samples can be compared on a scatterplot, where dot size corresponds to the number of clonotypes at a given coordinate.
Integrating gene expression data#
Leveraging the opportunity offered by close integeration with scanpy, transcriptomics-based data can be utilized alongside immune receptor data.
Clonotype modularity#
Using the Clonotype modularity we can identify clonotypes consisting of cells that are transcriptionally more similar than expected by random.
The clonotype modularity score represents the log2 fold change of the number of edges in the cell-cell neighborhood graph compared to the random background model. Clonotypes (or clonotype clusters) with a high modularity score consist of cells that have a similar molecular phenotype.
ir.tl.clonotype_modularity(mdata, target_col="airr:cc_aa_alignment")
Initalizing clonotype subgraphs...
Computing background distributions...
Stored result in `mdata.obs["airr:clonotype_modularity"]`.
Stored result in `mdata.obs["airr:clonotype_modularity_fdr"]`.
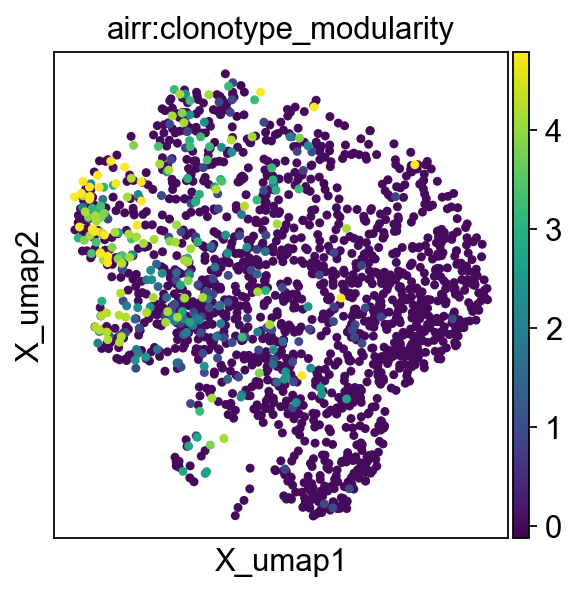
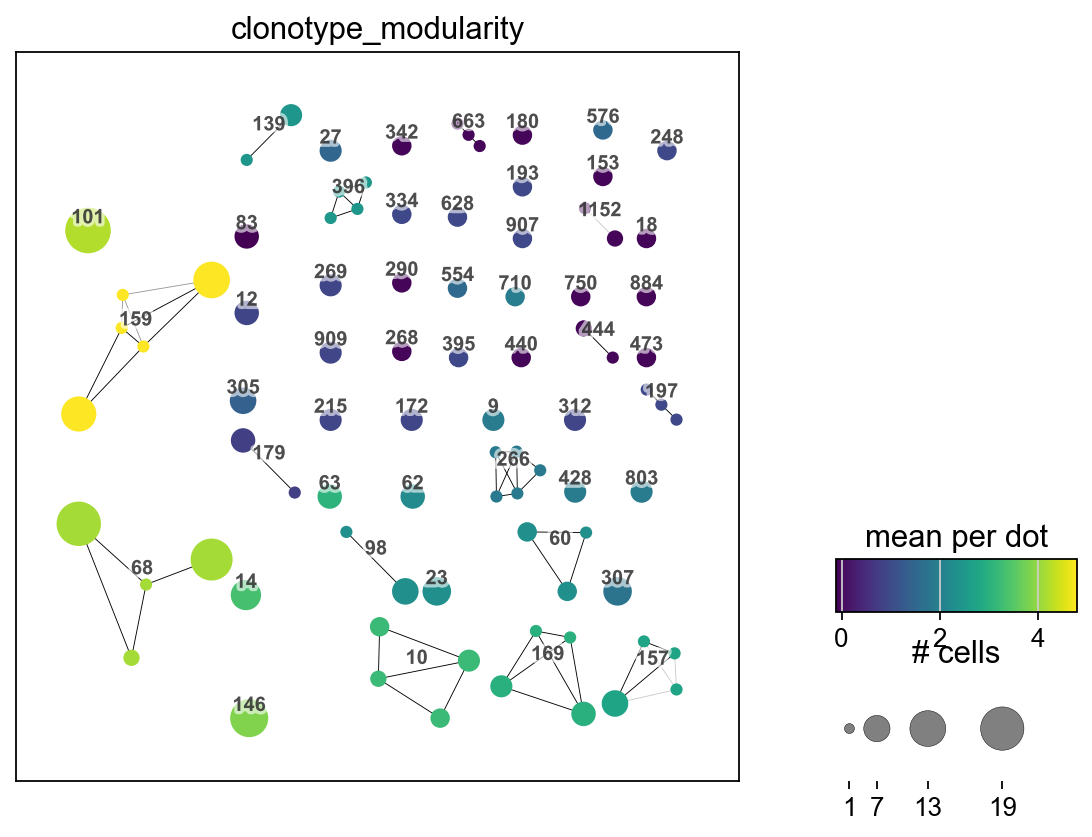
We can plot the clonotype modularity on top of a umap of clonotype network plot
_ = ir.pl.clonotype_network(
mdata,
color="clonotype_modularity",
label_fontsize=9,
panel_size=(6, 6),
base_size=20,
)
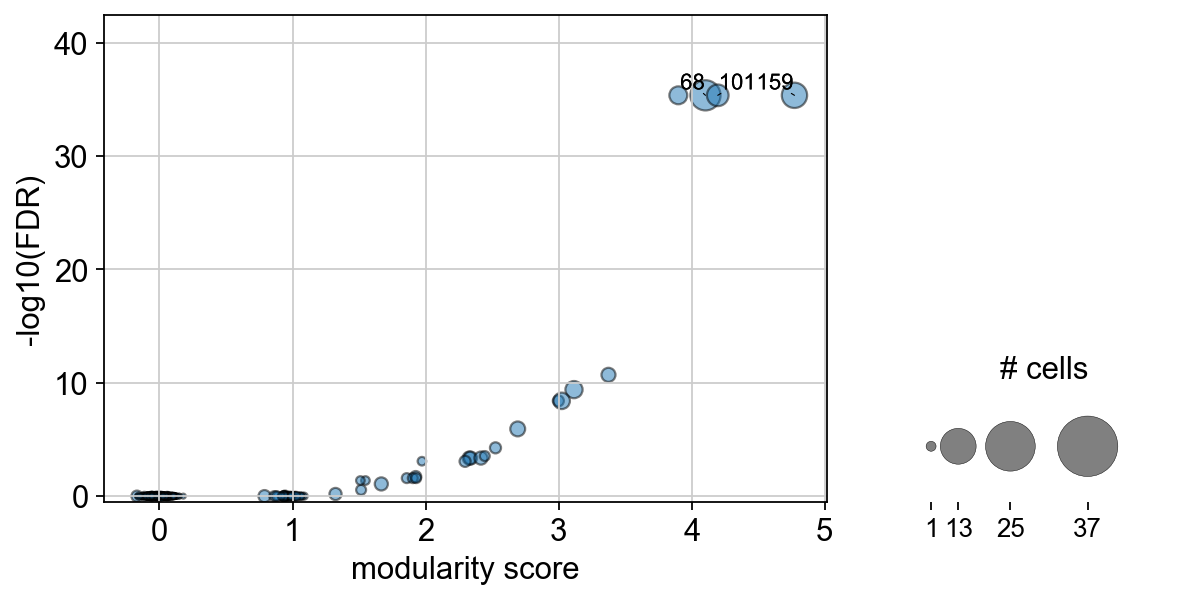
We can also visualize the clonotype modularity together with the associated FDR as a sort of “one sided volcano plot”:
ir.pl.clonotype_modularity(mdata, base_size=20)
Let’s further inspect the two top scoring candidates. We can extract that information from mdata.obs["airr:clonotype_modularity"].
clonotypes_top_modularity = list(
mdata.obs.set_index("airr:cc_aa_alignment")["airr:clonotype_modularity"]
.sort_values(ascending=False)
.index.unique()
.values[:2]
)
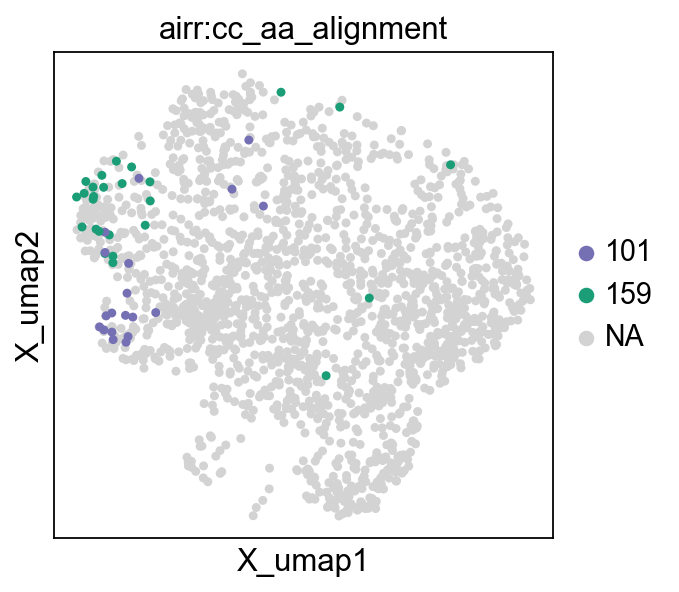
test_ad = mu.pl.embedding(
mdata,
basis="gex:umap",
color="airr:cc_aa_alignment",
groups=clonotypes_top_modularity,
palette=cycler(color=mpl_cm.Dark2_r.colors),
)
We observe that they are (mostly) restricted to a single cluster. By leveraging scanpy’s differential expression module, we can compare the gene expression of the cells in the two clonotypes to the rest.
# Since sc.tl.rank_genes_group does not support MuData, we need to temporarily add
# the AIRR columns to the gene expression AnnData object
with ir.get.obs_context(mdata["gex"], {"cc_aa_alignment": mdata.obs["airr:cc_aa_alignment"]}) as tmp_ad:
sc.tl.rank_genes_groups(
tmp_ad,
"cc_aa_alignment",
groups=clonotypes_top_modularity,
reference="rest",
method="wilcoxon",
)
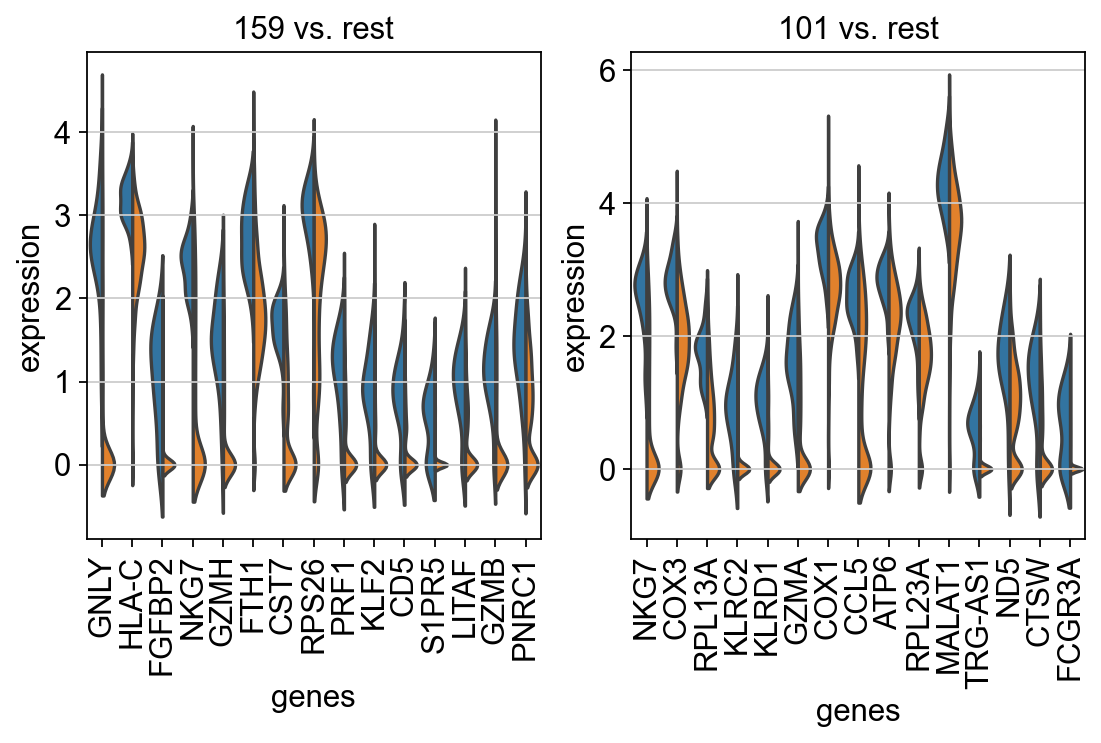
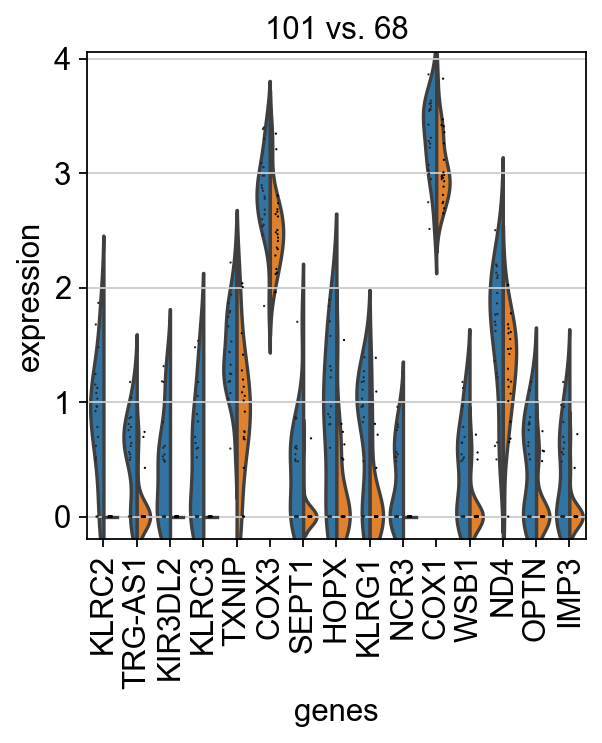
fig, axs = plt.subplots(1, 2, figsize=(8, 4))
for ct, ax in zip(clonotypes_top_modularity, axs):
sc.pl.rank_genes_groups_violin(tmp_ad, groups=[ct], n_genes=15, ax=ax, show=False, strip=False)
Clonotype imbalance among cell clusters#
Using cell type annotation inferred from gene expression clusters, for example, clonotypes belonging to CD8+ effector T-cells and CD8+ tissue-resident memory T cells, can be compared.
freq, stat = ir.tl.clonotype_imbalance(
mdata,
replicate_col="gex:sample",
groupby="gex:cluster",
case_label="CD8_Teff",
control_label="CD8_Trm",
inplace=False,
)
top_differential_clonotypes = stat["clone_id"].tolist()[:3]
WARNING: Clonotype imbalance calculation depends on repertoire overlap. We could not detect any previous runs of repertoire_overlap, so the tool is running now...
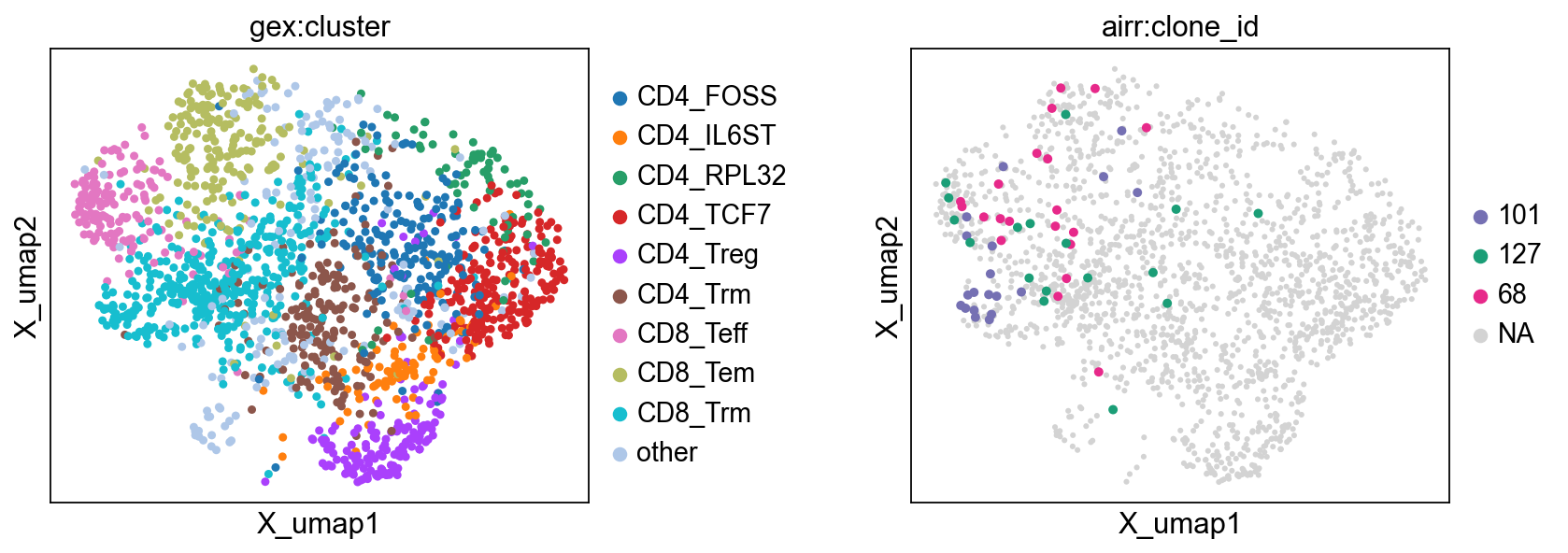
Showing top clonotypes on a UMAP clearly shows that clonotype 101 is featured by CD8+ tissue-resident memory T cells, while clonotype 68 by CD8+ effector and effector memory cells.
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 4), gridspec_kw={"wspace": 0.6})
mu.pl.embedding(mdata, basis="gex:umap", color="gex:cluster", ax=ax1, show=False)
mu.pl.embedding(
mdata,
basis="gex:umap",
color="airr:clone_id",
groups=top_differential_clonotypes,
ax=ax2,
# increase size of highlighted dots
size=[
80 if c in top_differential_clonotypes else 30 for c in mdata.obs["airr:clone_id"][mdata.mod["gex"].obs_names]
],
palette=cycler(color=mpl_cm.Dark2_r.colors),
)
Repertoire overlap of cell types#
Just like comparing repertoire overlap among samples, Scirpy also offers comparison between gene expression clusters or cell subpopulations. As an example, repertoire overlap of the two cell types compared above is shown.
Marker genes in top clonotypes#
Gene expression of cells belonging to individual clonotypes can also be compared using Scanpy. As an example, differential gene expression of two clonotypes, found to be specific to cell type clusters can also be analysed.
Query epitope databases#
We can use scirpy to query reference databases or datasets to annotate IRs with certain
features, such as epitope specificity. The reference database can be any dataset in scirpy’s AnnData format and you can follow the instructions in the data loading tutorial
to build a custom reference database, if it is not available from scirpy.datasets yet.
Querying reference datasets uses the same logic as defining clonotypes:
scirpy function |
objective |
|---|---|
Compute sequence-based distance matrices . |
|
For each cell, identify matching entries in a reference database. |
|
Transfer annotations from reference database to |
|
Return a dataframe with all matching annotations. |
Here, we obtain the VDJDB and annotate epitopes based
on amino acid sequence identity. For demonstration purposes on this toy dataset we use rather lenient settings: For a match, we specify that it is enough that either of the VJ and VDJ sequences, and either of the primary or secondary receptor chains matches the database.
vdjdb = ir.datasets.vdjdb()
Downloading latest version of VDJDB
Converting to AnnData object
Filtering chains...
Indexing VJ chains...
Indexing VDJ chains...
build result array
ir.pp.ir_dist(mdata, vdjdb, metric="identity", sequence="aa")
Computing sequence x sequence distance matrix for VJ sequences.
Computing sequence x sequence distance matrix for VDJ sequences.
ir.tl.ir_query(
mdata,
vdjdb,
metric="identity",
sequence="aa",
receptor_arms="any",
dual_ir="any",
)
Initializing lookup tables.
Computing clonotype x clonotype distances.
Stored IR distance matrix in `adata.uns["ir_query_VDJDB_aa_identity"]`.
ir_query_annotate_df() allows us to retrieve all pairs cells with their of matching entries. If a cell matches multiple entires from the reference database, the resulting data frame will contain multiple rows for the same cell.
ir.tl.ir_query_annotate_df(
mdata,
vdjdb,
metric="identity",
sequence="aa",
include_ref_cols=["antigen.species", "antigen.gene"],
).tail()
| antigen.species | antigen.gene | |
|---|---|---|
| RT3_TCTCTAATCACAATGC-1 | CMV | IE1 |
| RT3_TCTCTAATCACAATGC-1 | CMV | IE1 |
| RT3_TCTCTAATCACAATGC-1 | CMV | IE1 |
| RT3_TCTCTAATCACAATGC-1 | InfluenzaA | NP |
| RT3_TCTCTAATCACAATGC-1 | SARS-CoV-2 | ORF9b |
Alternatively, to break down the annotation to a single-value per cell, you can use
ir_query_annotate(). Depending on the specified strategy it will only label unambiguous matches, or use the most frequent value.
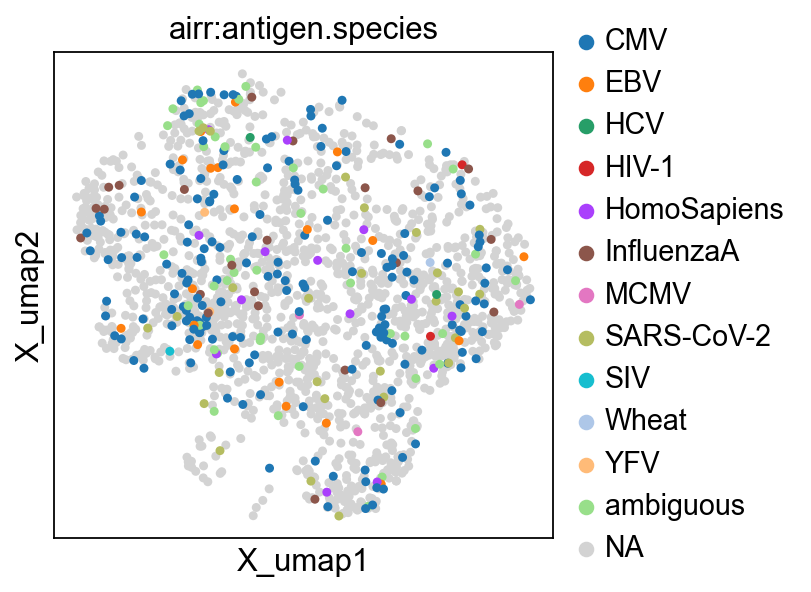
ir.tl.ir_query_annotate(
mdata,
vdjdb,
metric="identity",
sequence="aa",
include_ref_cols=["antigen.species"],
strategy="most-frequent",
)
Stored result in `mdata.obs["airr:antigen.species"]`.